13.8.6. Macro “reoptimizeZoneBiSubMpi.C”
13.8.6.1. Objective
The objective of the macro is to show an example of a two level parallelism program using the Mpi paradigm.
At the top level, an optimization loop parallelizes its evaluations
At low level, each optimizer evaluation are a launcher loop who parallelizes its own sequential evaluations
These example is inspired from a zoning problem of a small plant core with square assemblies. However, the physics embeded in it is reduced to none (sorry), and the problem is simplified. With symetries, the core is defined by 10 different assemblies presented on the following figure. For production purpose, only 5 assembly types are allowed, defined by an emission value.
Figure 13.59 The core and its assemblies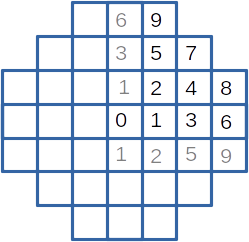
To simplify the problem, some constraints are put :
most assemblies belong to a default zone
other zone is restricted to one assembly (or two for 4 and 5, and for 8 and 9 for symetrical reason)
one zone is imposed with the 8th et 9th external assemblies.
the total of assembly emission is defined.
For each assembly, a reception value is defined depending on the emission from itself and its neighbour’s (just 8 neightbours are taken in account, the 4 nearest neighbours and 4 secondary neighbours). The global objective is to minimize the difference between the biggest and the smallest reception value.
Optimisation works on 4 emission values (the fifth value, affected to the external zone, is set, and all values are normalized with the total emission value) and each evaluation loops over the 35 possible arrangements (choose 3 zones from 7). A single evaluation take emission values and the selected zones and return the maximum reception difference.
13.8.6.2. Macro Uranie
This macro is splited in 2 files : the first one defines the low level evaluation function and is reused in the next reoptimizer example. It is quite a mock function, and is given to be complete, but is not needed to understand how to implement the two level MPI parallelism
/*
the different zones
6 9
3 5 7
1 2 4 8
0 1 3 6
1 2 5 9
*/
// 4 primary neighbours of a zone
int near1[10][4] = {
{1,1,1,1}, {0,2,2,3}, {1,1,4,5}, {1,4,5,6}, {2,3,7,8}, // 0-4
{2,3,7,9}, {3,8,9,10}, {4,5,10,10}, {4,6,10,10}, {5,6,10,10} // 5-9
};
// 4 secondary neighbours
int near2[10][4] = {
{2,2,2,2}, {1,1,4,5}, {0,3,3,7}, {2,2,8,9}, {1,5,6,10}, //4
{1,4,6,10}, {4,5,10,10}, {2,8,9,10}, {3,7,10,10}, {3,7,10,10} //9
};
// low level evaluation
void lowfun(double *in, double *out)
// evaluate a zoning
{
double dft;
double all, min, max, next;
int p, i, j, id;
double loc[11], bar[11];
// init dft
dft = in[0];
for (i=0; i<8; i++) loc[i] = dft;
loc[8] = loc[9] = 0.8;
loc[10] = 0.;
// init spec
for (i=4; i<7; i++) {
id = (int) in[i];
loc[id] = in[i-3];
if (id == 4) loc[5] = in[i-3];
}
// normalize
all = loc[0]/4;
for (i=1; i<11; i++) all += loc[i];
for (i=0; i<11; i++) loc[i] *= 10/all;
// max diff
i=0;
next = loc[i];
for (j=0; j<4; j++) next += loc[near1[i][j]]/8;
for (j=0; j<4; j++) next += loc[near2[i][j]]/16;
max = min = next;
for (i=1; i<10; i++) {
next = loc[i];
for (j=0; j<4; j++) next += loc[near1[i][j]]/8;
for (j=0; j<4; j++) next += loc[near2[i][j]]/16;
if (next < min) min = next;
else if (next > max) max = next;
}
out[0] = max-min;
for (i=0; i<10; i++) out[i+1] = loc[i];
}
The lowfun function deals, as expected, with the low level evaluation. In inputs it has the 4
emission values (default, zone1, zone2, zone3) and 3 indicators defining the zone affected by the
extra emission value. It returns the maximal difference between two zone reception values and the 9
normalized emission values (informative data). Two arrays are used to define the neighbourhood
With the second file, the two level MPI parallelism is defined.
using namespace URANIE::DataServer;
using namespace URANIE::Relauncher;
using namespace URANIE::Reoptimizer;
using namespace URANIE::MpiRelauncher;
#include "reoptimizeZoneCore.C"
void tds_resume(TDataServer *tds, TAttribute **att, double *res)
{
TList leaves;
TLeaf *leaf;
int i, j, k, siz;
double obj, cur;
std::vector<double> tmp;
siz = tds->getTuple()->GetEntries();
// init
for (i=0; att[i]; i++) {
leaves.Add(tds->GetTuple()->GetLeaf(att[i]->GetName()));
}
tmp.resize(i);
// search min
tds->GetTuple()->GetEntry(0);
obj = ((TLeaf *) leaves.At(0))->GetValue(0);
k = 0;
for (i=1; i<siz; i++) {
tds->GetTuple()->GetEntry(i);
cur = ((TLeaf *) leaves.At(0))->GetValue(0);
if (cur < obj) {
obj = cur;
k = i;
}
}
// get all results
TIter nextl(&leaves);
tds->GetTuple()->GetEntry(k);
for (j=0; (leaf = (TLeaf *) nextl() ); j++) {
res[j] = leaf->GetValue(0);
}
}
int doefun(double *in, double *out)
{
double z0, z1, z2, z3;
int i;
// const
z0 = in[0];
z1 = in[1];
z2 = in[2];
z3 = in[3];
// inputs
TAttribute zon0("zon0", 0., 1.);
TAttribute zon1("zon1", 0., 1.);
TAttribute zon2("zon2", 0., 1.);
TAttribute zon3("zon3", 0., 1.);
TAttribute a1("a1");
TAttribute a2("a2");
TAttribute a3("a3");
TAttribute *funi[] = { &zon0, &zon1, &zon2, &zon3, &a1, &a2, &a3, NULL};
//output
TAttribute diff("diff");
TAttribute v0("v0");
TAttribute v1("v1");
TAttribute v2("v2");
TAttribute v3("v3");
TAttribute v4("v4");
TAttribute v5("v5");
TAttribute v6("v6");
TAttribute v7("v7");
TAttribute v8("v8");
TAttribute v9("v9");
TAttribute *funo[] = {
&diff, &v0, &v1, &v2, &v3, &v4, &v5, &v6, &v7, &v8, &v9, NULL
};
// funlow
TCJitEval lfun(lowfun);
for (i=0; funi[i]; i++) lfun.addInput(funi[i]);
for (i=0; funo[i]; i++) lfun.addOutput(funo[i]);
// runner
// TSequentialRun run(&lfun);
TSubMpiRun run(&lfun);
run.startSlave();
if (run.onMaster()) {
TDataServer tds("doe", "tds4doe");
tds.keepFinalTuple(kFALSE);
for (i=4; i<7; i++) tds.addAttribute(funi[i]);
tds.fileDataRead("reoptimizeZoneDoe.dat", kFALSE, kTRUE, "quiet");
TLauncher2 launch(&tds, &run);
launch.addConstantValue(&zon0, z0);
launch.addConstantValue(&zon1, z1);
launch.addConstantValue(&zon2, z2);
launch.addConstantValue(&zon3, z3);
// run doe
launch.solverLoop();
//get critere
tds_resume(&tds, funo, out);
run.stopSlave();
}
return 1;
}
void reoptimizeZoneBiSubMpi()
{
//ROOT::EnableThreadSafety();
int i;
// inputs
TAttribute z1("zone1", 0., 1.);
TAttribute z2("zone2", 0., 1.);
TAttribute z3("zone3", 0., 1.);
TAttribute z4("zone4", 0., 1.);
TAttribute *zo[] = { &z1, &z2, &z3, &z4, NULL };
// outputs
TAttribute diff("diff");
TAttribute v0("v0");
TAttribute v1("v1");
TAttribute v2("v2");
TAttribute v3("v3");
TAttribute v4("v4");
TAttribute v5("v5");
TAttribute v6("v6");
TAttribute v7("v7");
TAttribute v8("v8");
TAttribute v9("v9");
TAttribute *out[] = {
&diff, &v0, &v1, &v2, &v3, &v4, &v5, &v6, &v7, &v8, &v9, NULL
};
// fonction
TCJitEval fun(doefun);
for (i=0; zo[i]; i++) fun.addInput(zo[i]);
for (i=0; out[i]; i++) fun.addOutput(out[i]);
fun.setMpi();
// runner
//TThreadedRun runner(&fun,8);
//TSequentialRun runner(&fun);
TBiMpiRun runner(&fun, 3);
runner.startSlave();
if (runner.onMaster()) {
TDataServer tds("tdsvzr", "tds4optim");
fun.addAllInputs(&tds);
//
TVizirGenetic gene;
gene.setSize(300, 200000, 100);
//TVizirIsland viz(&tds, &runner, &gene);
TVizir2 viz(&tds, &runner, &gene);
//viz.setTolerance(0.00001);
viz.addObjective(&diff);
viz.solverLoop();
runner.stopSlave();
tds.exportData("__coeur__.dat");
}
}
This script is structured with 3 functions :
function
tds_resumeis used by the intermediate function. It receives theTDataServerfilled, loops on its items and returns an synthetic value. In our case, the minimum value of the reception difference, and the 9 normalized emission valuesfunction
doefunis the intermediate evaluation function. It runs the design of experiments containing all 35 possible arrangements and extract the best one. It receives the 4 emission values and used them to complete theTDataServerusing theaddConstantValuemethod.function
reoptimizeZoneBiSubMpiis the top level function who solve the zoning problem
TBiMpiRun and TSubMpiRun are used to allocate cpus between intermediate and low level.
TBiMpiRun is used in reoptimizeZoneBiSubMpi (top) with an integer argument specifying the
number of CPUs dedicated to each intermediate level. In our case (3), with 16 resources request to
MPI, they are divided in 5 groups of 3 CPUs, and one CPU is left for the top level master (take care
that the number of CPUs requested matches group size (16 % 3 == 1)). The top level Master sees 5
resources for his evaluations. TSubMpiRun is used in doefun function and gives access to the 3
own resources reserved in top level function.
Running the script is done as usual with MPI :
mpirun -n 16 root -l -b -q reoptimizeZoneBiSubMpi.C
At the begining of reoptimizeZoneBiSubMpi function there is a call to ROOT::EnableThreadSafety.
It is unusefull in this case, but if we parallelize with threads instead of MPI. If you want to use
both threads and MPI, it is recommended to use MPI at top level.