13.9.5. Macro “reoptimizeHollowBarVizirSplitRuns.py”
13.9.5.1. Objective
The objective of the macro is to be able to run an evolutionary algorithm (here we are using a genetic one) with a limited number of code estimation and restart it from where it stopped if it has not converged the first time. This is of utmost usefulness when running a resource-consumming code or (/and) when running on a cluster with a limited number of cpu time. The classical hollow bar example defined in Sizing of a hollow bar example problem is used to obtain a nice Pareto set/front.
13.9.5.2. Macro Uranie
"""
Example of hollow bar optimisation in split runs
"""
from URANIE import DataServer, Relauncher, Reoptimizer
import ROOT
TOLERANCE = 0.001
NBmaxEVAL = 1200
SIZE = 500
def launch_vizir(run_number, fig_one):
"""Factorised function to run the Vizir analaysis."""
# variables
xvar = DataServer.TAttribute("x", 0.0, 1.0)
yvar = DataServer.TAttribute("y", 0.0, 1.0)
thick = DataServer.TAttribute("thick")
sect = DataServer.TAttribute("sect")
defor = DataServer.TAttribute("defor")
code = Relauncher.TCIntEval("barAllCost")
code.addInput(xvar)
code.addInput(yvar)
code.addOutput(thick)
code.addOutput(sect)
code.addOutput(defor)
# Create a runner
runner = Relauncher.TSequentialRun(code)
runner.startSlave()
# Output to state whether convergence is reached
has_converged = False
if runner.onMaster():
# Create the TDS
tds = DataServer.TDataServer("vizirDemo", "Param de l'opt vizir")
tds.addAttribute(xvar)
tds.addAttribute(yvar)
solv = Reoptimizer.TVizirGenetic()
# Name of the file that will contain
filename = "genetic.dump"
# whether = Test if a genetic.dump exists. If not, it creates it
# and returns false, so that the "else" part is done to start the
# initialisation of the vizir algorithm.
if solv.setResume(NBmaxEVAL, filename):
print("Restarting Vizir")
else:
solv.setSize(SIZE, NBmaxEVAL)
# Create the multi-objective constrained optimizer
opt = Reoptimizer.TVizir2(tds, runner, solv)
opt.setTolerance(TOLERANCE)
# add the objective
opt.addObjective(sect)
opt.addObjective(defor)
positiv = Reoptimizer.TGreaterFit(0.4)
opt.addConstraint(thick, positiv)
# resolution
opt.solverLoop()
has_converged = opt.isConverged()
# Stop the slave processes
runner.stopSlave()
fig_one.cd(run_number+1)
tds.getTuple().SetMarkerColor(2)
tds.draw("defor:sect")
tit = "Run number "+str(run_number+1)
if has_converged:
tit += ": Converged !"
ROOT.gPad.GetPrimitive("__tdshisto__0").SetTitle(tit)
return has_converged
ROOT.gROOT.LoadMacro("UserFunctions.C")
# Delete previous file if it exists
ROOT.gSystem.Unlink("genetic.dump")
finished = False
i = 0
fig1 = ROOT.TCanvas("fig1", "fig1", 1200, 800)
fig1.Divide(2, 2)
while not finished:
finished = launch_vizir(i, fig1)
i = i+1
The idea is to show how to run this kind of configuration: the function LaunchVizir is the usual
script one can run to get an optimisation with Vizir on the hollow bar problem. The aim is to create
a Pareto set of 500 points (SIZE) but only allowing 1200 estimation (NBmaxEVAL). With this
configuration we are sure that a first round of estimation will not converge, so we will have to
restart the optimisation from the point we stopped. With this regard, the beginning of this function
is trivial and the main point to be discussed arises once the solver is created.
solv = Reoptimizer.TVizirGenetic()
# Name of the file that will contain
filename = "genetic.dump"
# whether = Test if a genetic.dump exists. If not, it creates it
# and returns false, so that the "else" part is done to start the
# initialisation of the vizir algorithm.
if solv.setResume(NBmaxEVAL, filename):
print("Restarting Vizir")
else:
solv.setSize(SIZE, NBmaxEVAL)
Clearly here, the interesting part apart, from the definition of the name of the file in which the
final state will be kept, is the first test on the solver, before using the setSize method. A new
methods called setResume is called, with two arguments : the number of elements requested in the
Pareto set and the name of the file in which to save the state or to restart from. This method
returns “true” if genetic.dump is found and “false” if not. In the first case, the code will assume
that this file is the result of a previous run and it will start the optimisation from the its
content trying to get all the population non-dominated (if it’s not yet the case). If, on the other
hand, no file is found, then the code knows that it would have to store the results of its process,
in a file whose name is the second argument, and because the function returns “false”, then we move
to the “else” part, that starts the optimisation.
Apart from this, the rest of the function is doing the optimisation, and plotting the pareto front in a provided canvas. The only new part here is the fact that the solver (its master in fact) is now able to tell whether it has converged or not through the following method
has_converged = opt.isConverged()
this argument being return as the results of the function.
The rest of this macro plays the role of the user in front of a ROOT-console (its python interplay of course).
It defines the correct namespace, loads the function file and destroys previously existing
genetic.dump files. From there it runs the LaunchVizir function as many times as needed (thanks
to the boolean returned) as the used would do, by restarting the macro, even after exiting the ROOT
console.
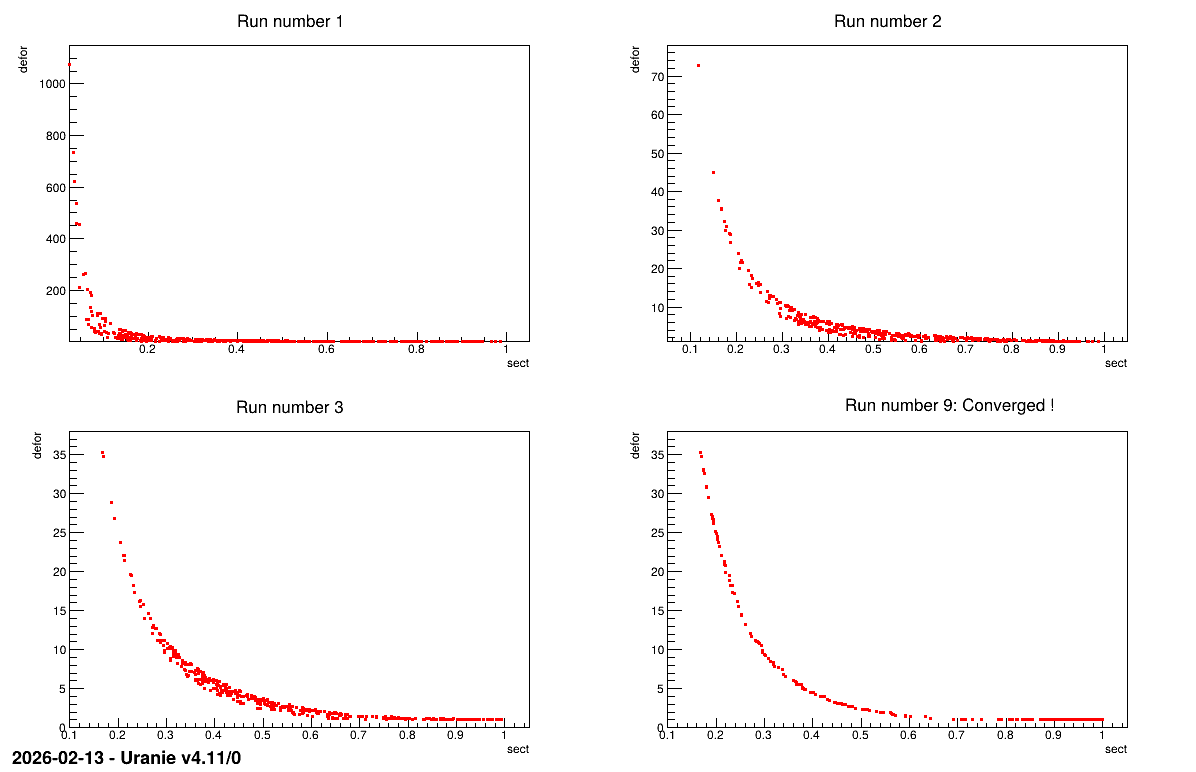
The plot shown below represent the Pareto front every time the genetic algorithm stops (at the fourth run, it finally converges !).
13.9.5.3. Graph
Figure 13.57 Graph of the macro “reoptimizeHollowBarVizirSplitRuns.py”