13.10.5. Macro “calibrationMCMCFlowrate1D.C”
13.10.5.1. Objective
The goal here is to calibrate the parameter \(H_l\) within the flowrate model, while varying only two
inputs (\(r_{\omega}\) and \(L\)). The remaining variables are fixed to the following values: \(r=25050\),
\(T_u=89335\), \(T_l=89.55\), \(H_u=1050\), \(K_{\omega}=10950\). The context of this example has already been
presented in Use-case for this chapter, including the model (implemented
here as a C++ function) and the initial lines defining the TDataServer objects. This macro demonstrates how
to apply the Markov chain Monte Carlo approach and more precisely the component-wise Metropolis-Hastings algorithm using a Relauncher-based architecture.
13.10.5.2. Macro Uranie
TDataServer *tdsRef = new TDataServer("tdsRef","doe_exp_Re_Pr");
TDataServer *tdsPar = new TDataServer("tdsPar","tdsPar");
// Load the function flowrateCalib1D
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="Ex2DoE_n100_sd1.75.dat";
// Define the reference
tdsRef->fileDataRead(ExpData.Data());
// Define the parameters
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib1D");
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
// Set the calibration object
TMCMC *cal = new TMCMC(tdsPar, runner, 2000);
// Set the likelihood and reference with default gaussian log likelihood
cal->setLikelihood("log-gauss",tdsRef,"rw:l","Qexp","sd_eps");
// Set the MCMC algorithm to component-wise Metropolis-Hastings
cal->setAlgo("MH_1D");
// Set the acceptation ratio range between 20% and 50%
cal->setAcceptationRatioRange(0.2, 0.5);
// Set the number of iterations between two information displays
cal->setNbDump(500);
// Initialise 4 chains
cal->setMultistart(4);
// Set the starting points of each chain
cal->setStartingPoints(0, {740});
cal->setStartingPoints(1, {745});
cal->setStartingPoints(2, {750});
cal->setStartingPoints(3, {755});
// Set the standard deviation of the proposal for all the chains
cal->setProposalStd(-1, {5});
// Run the MCMC algorithm
cal->estimateParameters();
// Quality assessment: Draw the trace of the Markov Chain
TCanvas *canTr = new TCanvas("CanTr","CanTr",1200,800);
TPad *padTr = new TPad("padTr","padTr",0, 0.03, 1, 1);
padTr->Draw();
padTr->cd();
cal->drawTrace("Trace", "*", "nonewcanvas");
// Quality assessment: Draw the acceptation ratio
TCanvas *canAcc = new TCanvas("CanAcc","CanAcc",1200,800);
TPad *padAcc = new TPad("padAcc","padAcc",0, 0.03, 1, 1);
padAcc->Draw();
padAcc->cd();
cal->drawAcceptationRatio("Acceptation ratio", "*", "nonewcanvas");
// Set the number of iterations to remove
int burnin = 50;
cal->setBurnin(burnin);
// Verify that the burn-in has been set correctely with Gelman-Rubin statistic
std::unordered_map<string, double> GelmanRubin_values = cal->diagGelmanRubin();
// Compute Effective Sample Size
std::unordered_map<string, vector<int>> ESS_values = cal->diagESS();
// Set the lag
int lag = 1;
cal->setLag(lag);
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals", "*", "", "nonewcanvas");
// Draw the posterior distributions
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1);
padPar->Draw();
padPar->cd();
cal->drawParameters("Posterior", "*", "nonewcanvas");
Much of this code has already been covered in the previous section
Macro “calibrationMinimisationFlowrate1D.C” (up to the sequential run). The main difference here
is that the input parameter is now defined as a TStochasticDistribution, representing the a priori
chosen distribution.
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
The calibration object is then created by specifying both the number of iterations of the algorithm (set to 2000). The likelihood function is defined as Gaussian, with the reference dataserver, the input and output variables, and the standard deviation of the experimental data provided as arguments. Several additional options are configured: the component-wise algorithm is selected, the acceptance range is set, the interval between two console displays is specified, the number of chains to initialise is defined, the starting points of each chain are assigned (one by one), and the standard deviation of the initial proposal distribution is set (for all chains at once). For further details on these options, see Defining the TMCMC properties. Finally, the estimation process is performed, using the chosen MCMC algorithm (component-wise Metropolis-Hastings).
// Set the calibration object
TMCMC *cal = new TMCMC(tdsPar, runner, 2000);
// Set the likelihood and reference with default gaussian log likelihood
cal->setLikelihood("log-gauss",tdsRef,"rw:l","Qexp","sd_eps");
// Set the MCMC algorithm to component-wise Metropolis-Hastings
cal->setAlgo("MH_1D");
// Set the acceptation ratio range between 20% and 50%
cal->setAcceptationRatioRange(0.2, 0.5);
// Set the number of iterations between two information displays
cal->setNbDump(500);
// Initialise 4 chains
cal->setMultistart(4);
// Set the starting points of each chain
cal->setStartingPoints(0, {740});
cal->setStartingPoints(1, {745});
cal->setStartingPoints(2, {750});
cal->setStartingPoints(3, {755});
// Set the standard deviation of the proposal for all the chains
cal->setProposalStd(-1, {5});
// Run the MCMC algorithm
cal->estimateParameters();
The final part demonstrates how to display and analyse the results. The first elements to examine are the trace plot (Figure 13.67) and the acceptance ratio plot (Figure 13.68). On these plots, you should check that the number of accepted iterations is sufficiently high (at least several hundred if the chains stabilise quickly — in our case, around 500), and that the converged acceptance ratio falls within a reasonable range (20–50% — here, it is close to 25%). When inspecting the trace, the chains should have converged around the same region (as they do here), and they should evolve rapidly, which indicates efficient exploration and low autocorrelation (also observed in our case). Finally, these plots help verify whether all chains behave stably and from which iteration they can be considered stable. This provides a first indication of a suitable burn-in value. In this example, the chains appear to have converged, and a burn-in value of 50 seems appropriate.
The burn-in size can then be defined and its consistency verified using the Gelman–Rubin statistic. In our case, a value of 1.00137 is reported in Console which is very close to 1 and confirms excellent convergence of the chains. Otherwise, a different burn-in value might have been more appropriate, the calculation could have been extended, or the MCMC algorithm settings adjusted.
The next step is to check for sample autocorrelation within the chains. The Effective Sample Size (ESS) provides a good indication of the lag to be used. In our case, the chains contain a reasonable number of independent samples (at least 257 each), and a lag of 1 is sufficient (see Console). Otherwise, the calculation could have been extended, or the MCMC algorithm settings adjusted (especially the standard deviation of the proposal distribution).
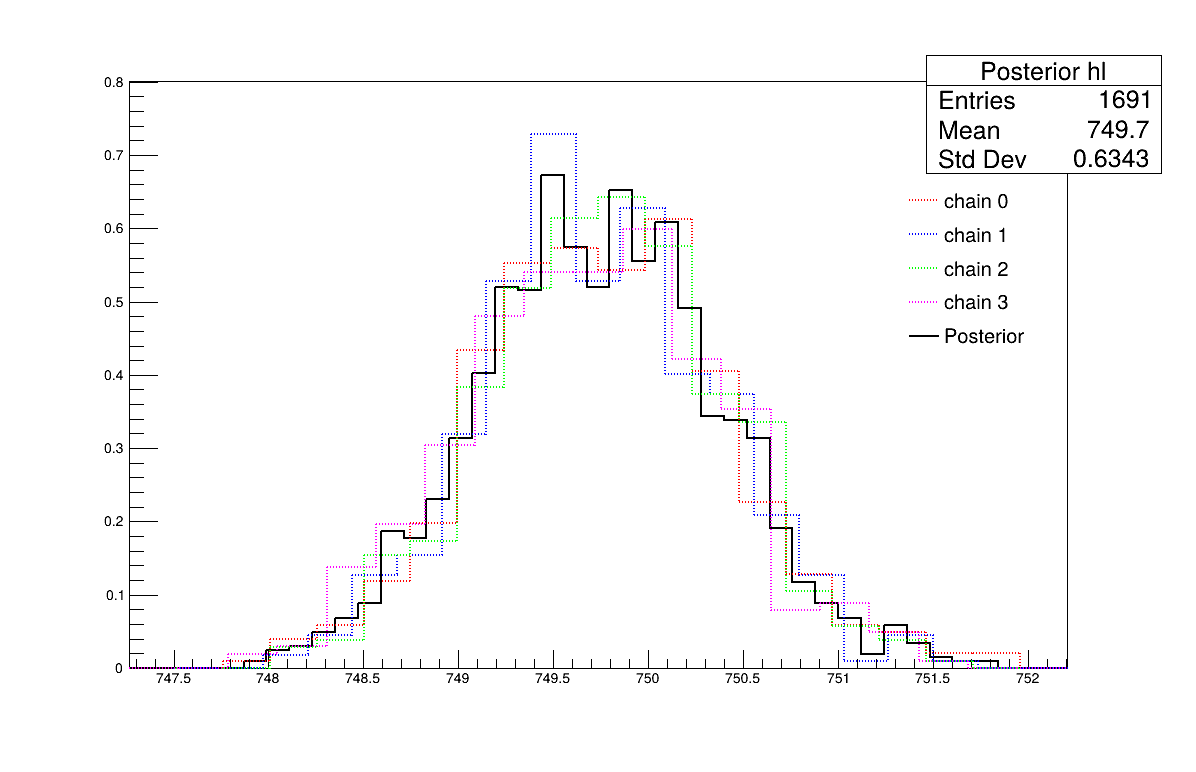
Two additional pieces of a posteriori information are presented as plots: the residuals (Figure 13.69), which show the expected normal distribution behavior, as discussed in [Bla17] and the posterior distribution (Figure 13.70).
// Quality assessment: Draw the trace of the Markov Chain
TCanvas *canTr = new TCanvas("CanTr","CanTr",1200,800);
TPad *padTr = new TPad("padTr","padTr",0, 0.03, 1, 1);
padTr->Draw();
padTr->cd();
cal->drawTrace("Trace", "*", "nonewcanvas");
// Quality assessment: Draw the acceptation ratio
TCanvas *canAcc = new TCanvas("CanAcc","CanAcc",1200,800);
TPad *padAcc = new TPad("padAcc","padAcc",0, 0.03, 1, 1);
padAcc->Draw();
padAcc->cd();
cal->drawAcceptationRatio("Acceptation ratio", "*", "nonewcanvas");
// Set the number of iterations to remove
int burnin = 50;
cal->setBurnin(burnin);
// Verify that the burn-in has been set correctely with Gelman-Rubin statistic
std::unordered_map<string, double> GelmanRubin_values = cal->diagGelmanRubin();
// Compute Effective Sample Size
std::unordered_map<string, vector<int>> ESS_values = cal->diagESS();
// Set the lag
int lag = 1;
cal->setLag(lag);
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals", "*", "", "nonewcanvas");
// Draw the posterior distributions
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1);
padPar->Draw();
padPar->cd();
cal->drawParameters("Posterior", "*", "nonewcanvas");
13.10.5.3. Console
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Thu Feb 12, 2026
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[/volatile/projet/uranie-ci/builds/Mfxi5pax-/0/uranie-cea/uranie/souRCE/meTIER/calibration/souRCE/TMCMC.cxx] Line[193]
<URANIE::INFO> TMCMC::constructor
<URANIE::INFO> A folder named [MCMC_1] has been created to store the results of the new calculation.
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[/volatile/projet/uranie-ci/builds/Mfxi5pax-/0/uranie-cea/uranie/souRCE/meTIER/calibration/souRCE/TMCMC.cxx] Line[673]
<URANIE::INFO> TMCMC::constructor
<URANIE::INFO> MCMC_1_chain_0 file has been duplicated in the folder [MCMC_1] to initiate [4] chains.
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [22.95%].
A posteriori mean coordinates are : (749.692)
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [25.8%].
A posteriori mean coordinates are : (749.727)
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [23.55%].
A posteriori mean coordinates are : (749.769)
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [22.05%].
A posteriori mean coordinates are : (749.713)
Gelman-Rubin statistic for variable [hl] is equal to 1.00137. Excellent convergence.
Effective Sample Size (ESS) for variable [hl] and chain [0] is equal to 257.
Effective Sample Size (ESS) for variable [hl] and chain [1] is equal to 305.
Effective Sample Size (ESS) for variable [hl] and chain [2] is equal to 405.
Effective Sample Size (ESS) for variable [hl] and chain [3] is equal to 371.
For uncorrelated samples, the lag should be set to 1.
13.10.5.4. Graphs
Figure 13.67 Trace graph of the macro “calibrationMCMCFlowrate1D.C”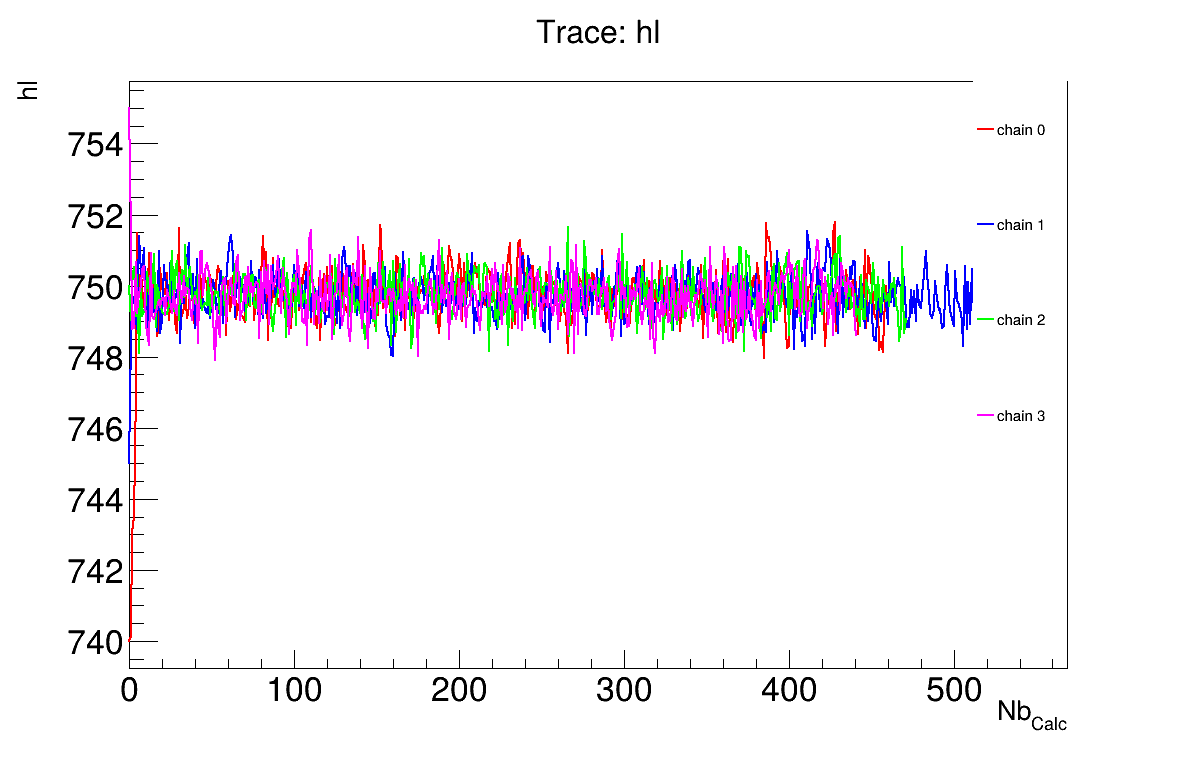
Figure 13.68 Acceptation ratio graph of the macro “calibrationMCMCFlowrate1D.C”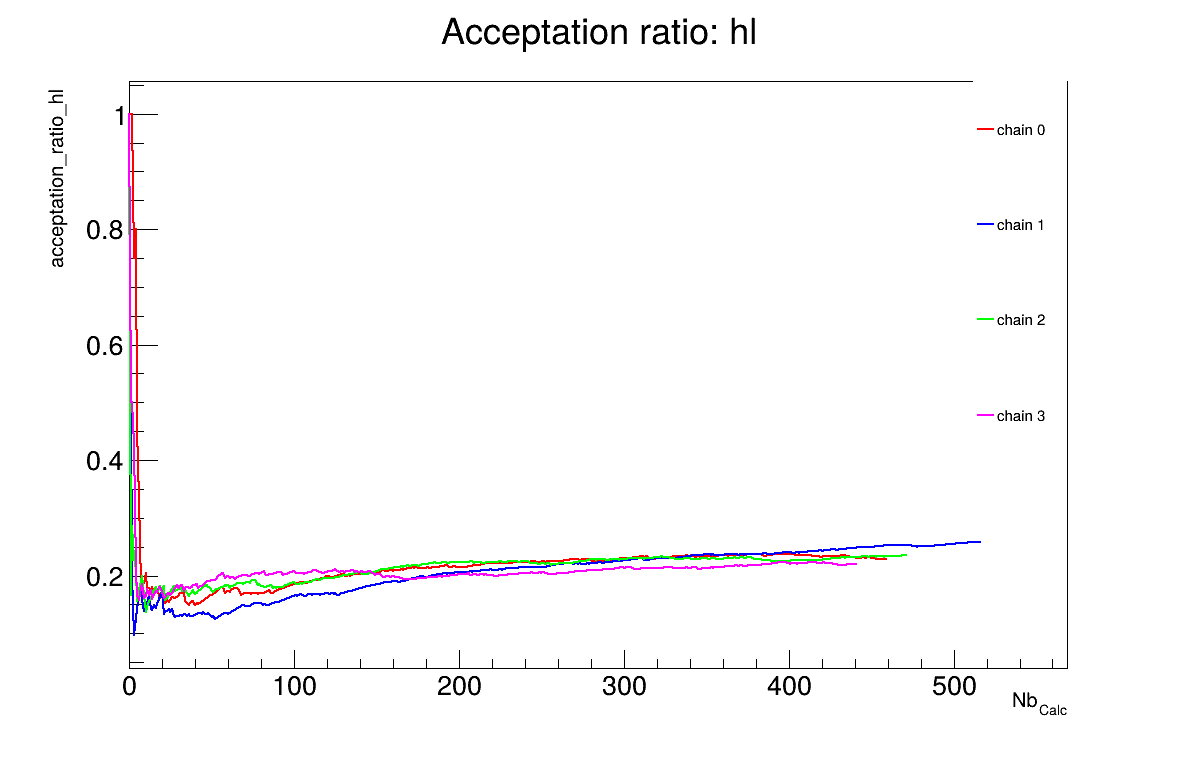
Figure 13.69 Residuals graph of the macro “calibrationMCMCFlowrate1D.C”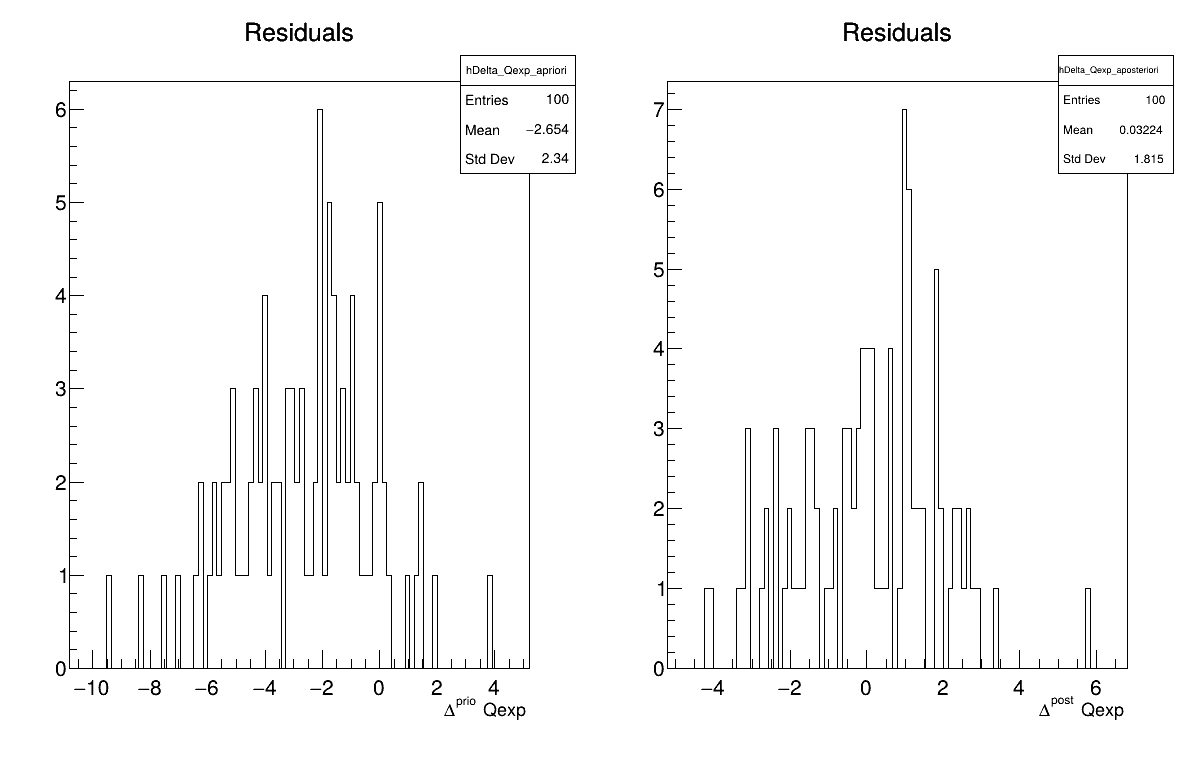
Figure 13.70 Parameter graph of the macro “calibrationMCMCFlowrate1D.C”