13.4.13. Macro “samplingConstrLHSEllipses.py”
13.4.13.1. Objective
This macro shows the usage of the TConstrLHS class when one wants to create a constrained LHS with
non-linear constraints. In order to illustate the concept, it is applied on three input variables
drawn from uniform distribution with well-thought boundaries (as the concept of the LHS is to have
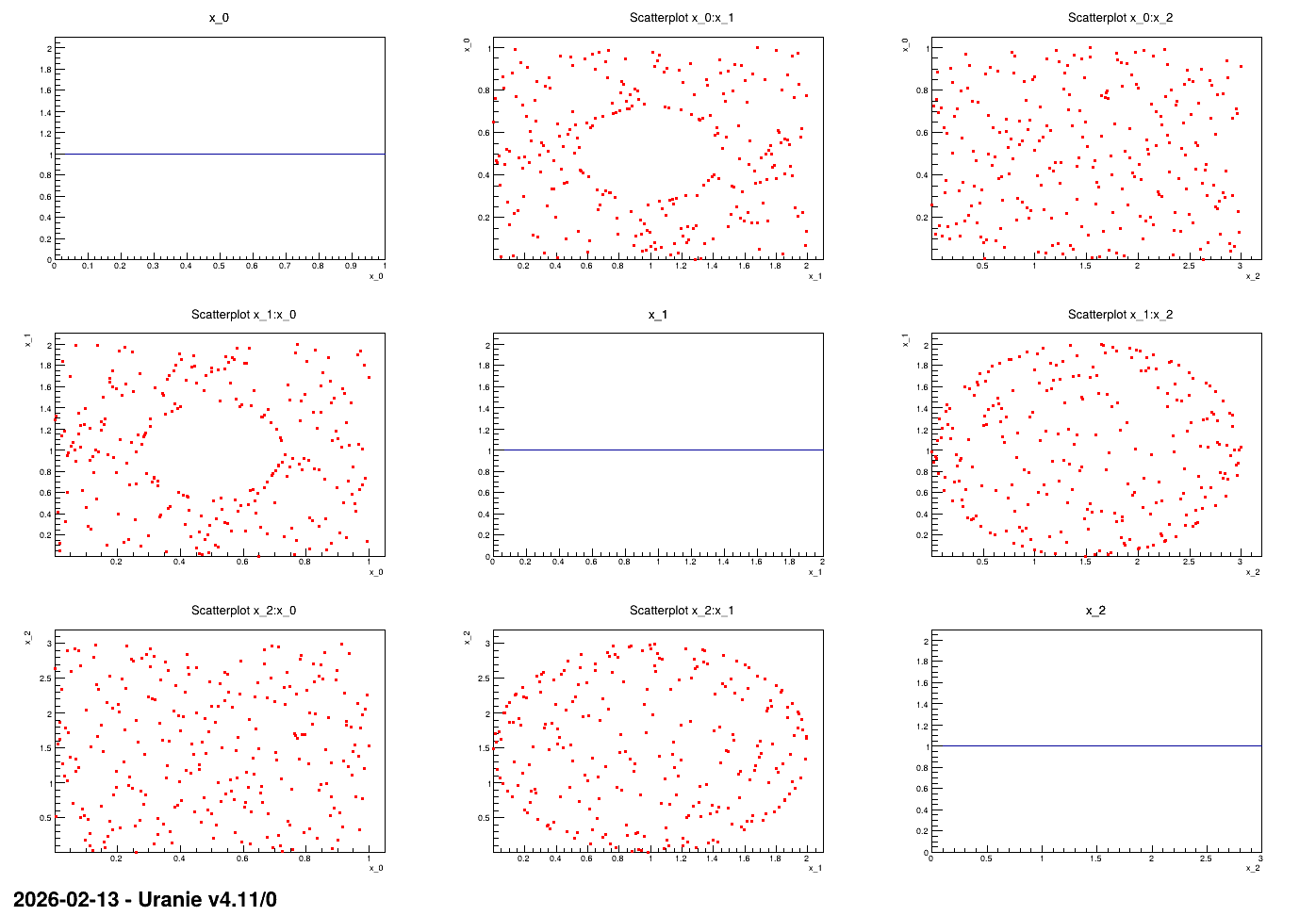
nicely distributed marginals). The constraints are excluding the inner part of an ellipse for one of
the input plane and the outter part of another ellipse for another input plane.
13.4.13.2. Macro Uranie
"""
Example of Constrained lhs with ellipses
"""
import numpy as np
from URANIE import DataServer, Sampler
import ROOT
def nicegrid(l_tds, l_can):
"""Generate the grid plot."""
l_ns = l_tds.getNPatterns()
l_nx = l_tds.getNAttributes()
l_tds.drawPairs()
all_h = []
ROOT.gStyle.SetOptStat(0)
for iatt in range(l_nx):
l_can.GetPad(iatt*(l_nx+1)+1).cd()
att = l_tds.getAttribute(iatt)
a_name = att.GetName()
all_h.append(ROOT.TH1F(a_name+"_histo", a_name+";x_"+str(iatt),
l_ns, att.getLowerBound(), att.getUpperBound()))
l_tds.Draw(a_name+">>"+a_name+"_histo", "", "goff")
all_h[iatt].Draw()
return all_h
# Canvas to produce the pair plot
Can = ROOT.TCanvas("Can", "Can", 10, 32, 1400, 1000)
apad = ROOT.TPad("apad", "apad", 0, 0.03, 1, 1)
apad.Draw()
apad.cd()
ns = 250 # Size of the samples to be produced
nx = 3
# Create dataserver and define the attributes
tds = DataServer.TDataServer("tds", "pouet")
for i in range(nx):
tds.addAttribute(DataServer.TUniformDistribution("x_"+str(i), 0, i+1))
tds.getAttribute("x_"+str(i)).delShare()
ROOT.gROOT.LoadMacro("ConstrFunctions.C")
# Generate the constr lhs
constrlhs = Sampler.TConstrLHS(tds, ns)
inputs = np.array([1, 0, 1, 2], dtype='i')
constrlhs.addConstraint(ROOT.CircularRules, 2, len(inputs), inputs)
constrlhs.generateSample()
# Do the plot
ListOfDiag = nicegrid(tds, apad)
The very beginning of these macros is the nicegrid method which is here only to show the nice
marginal distributions and the scatter plots. One can clearly skip this part to focus on the rest in
the main function.
The macro very much looks like any other design-of-experiments generating macro above: the dataserver is created
along with the canvas object and the problem is defined along with the input variables and the number
of locations to be produced. Once done, then the TConstrLHS instance is created with the four
following lines:
# Generate the constr lhs
constrlhs = Sampler.TConstrLHS(tds, ns)
inputs = np.array([1, 0, 1, 2], dtype='i')
constrlhs.addConstraint(ROOT.CircularRules, 2, len(inputs), inputs)
constrlhs.generateSample()
The constructor is pretty obvious, as it takes only the dataserver object and the number of locations.
Once created the main method to be called is the addConstraint function which has been largely
discussed in TConstrLHS example. The first argument of this method is the
pointer to the C++ function which has been included in our macro through the line used right before the sampler creation:
ROOT.gROOT.LoadMacro("ConstrFunctions.C")
which contains the CircularRules function. The rest of the argument are the number of constraints,
the size of the list of parameters and its content. Finally the nicegrid method is called to
produce the nice plot shown in Figure 13.16
13.4.13.3. Graph
Figure 13.16 Graph of the macro “samplingConstrLHSEllipses.py”