13.3.14. Macro “samplingSingularCorrelationCase.C”
13.3.14.1. Objective
This macro shows the usage of the SVD decomposition for the specific case where the target correlation
matrix is singular. The idea is to provide a tool that will allow the user to compare quickly both
the TSampling and TBasicSampling implementation, with a singular correlation matrix, or not. In
order to do that, a toy random correlation matrix generator is provided.
The resulting design-of-experiments is presented side-by-side with the residual of the obtained correlation matrix.
13.3.14.2. Macro Uranie
// Creating the TDS
TDataServer *tds = new TDataServer("pouet","pouet");
// What classes to be used (true==TSampling, false==TBasicSampling)
bool ImanConover=false;
TString SamplingType="srs";
TString SamplerOption="svd"; // Remove svd to use Cholesky
// Generating randomly a singular correlation matrix
// dimension of the problem in total.
int n = 10;
// number of dimension self explained.
// SHOULD BE SMALLER (singular) OR EQUAL TO (full-rank) n.
int p = 6;
// number of location in the doe
int m=300;
// =========================================================================
// Internal recipe to get this correlation matrix
// =========================================================================
TMatrixD A(n,p);
TRandom3 *Rand = new TRandom3();
for(int i=0;i<n;i++){
for(int j=0;j<p;j++){
A(i,j) = Rand->Gaus(0.,1.);
}
}
TMatrixD Gamma(A,TMatrixD::kMultTranspose,A);
Gamma*=1./n;
TMatrixD Sig(n,n);
for(int i=0;i<n;i++) Sig(i,i)=1./sqrt( Gamma(i,i) );
TMatrixD Corr(Sig,TMatrixD::kMult, Gamma);
Corr*=Sig;
// =========================================================================
// Corr is our correlation matrix.
// =========================================================================
// Adding attributes
for(int i=0;i<n;i++) tds->addAttribute( new TNormalDistribution(Form("n%d",i),0.,1.) );
// Create the sampler and generate the doe
if(ImanConover){
TSampling *sam = new TSampling(tds,SamplingType.Data(),m);
sam->setCorrelationMatrix(Corr);
sam->generateSample(SamplerOption.Data());
}else{
TBasicSampling *sam = new TBasicSampling(tds,SamplingType.Data(),m);
sam->setCorrelationMatrix(Corr);
sam->generateCorrSample(SamplerOption.Data());
}
// Compute the empirical correlation matrix
TMatrixD ResultCorr=tds->computeCorrelationMatrix();
// Change it into a residual matrix
ResultCorr-=Corr;
gStyle->SetOptStat(1110);
// Plot the results
TCanvas *c=new TCanvas("c","c",1800,900);
TPad *apad=new TPad("apad","apad",0, 0.03, 1, 1); apad->Draw();
apad->cd();
apad->Divide(2,1);
apad->cd(1);
// Residual distribution
TH1F *h=new TH1F("h",";#Delta_{#rho}",100,-0.2,0.2);
for(int i=0;i<n;i++)
{
for(int j=i;j<n;j++) h->Fill(ResultCorr(i,j));
}
h->Draw();
// all variables
apad->cd(2);
tds->drawPairs();
This macro starts by a bunch of variables provided to offer many possible configuration, among which:
use Iman and Conover method or the more simple one to get the correlation (see [Bla17] for a complete description of their respective correlation handling);
produce a stratified or fully random sample;
use Cholesky or SVD decomposition to decompose the target correlation matrix;
use a full-rank or singular correlation matrix. This is allowed thanks to the toy random correlation matrix generator: one has to define the number of variable (n) and the number of dimension that should be self-explained (p). If the latter is strickly lower than the former, then the correlation matrix generated will be singular, whereas it will be a full-rank one if both quantities are equal.
This is the main part of this macro
// What classes to be used (true==TSampling, false==TBasicSampling)
bool ImanConover=false;
TString SamplingType="srs";
TString SamplerOption="svd"; // Remove svd to use Cholesky
// Generating randomly a singular correlation matrix
// dimension of the problem in total.
int n = 10;
// number of dimension self explained.
// SHOULD BE SMALLER (singular) OR EQUAL TO (full-rank) n.
int p = 6;
// number of location in the doe
int m=300;
Once this is settled, the correlation matrix is created and the dataserver is created and n centered-reduced normal attributes are added. The chosen design-of-experiments is generated with the chosen options:
// Create the sampler and generate the doe
if(ImanConover){
TSampling *sam = new TSampling(tds,SamplingType.Data(),m);
sam->setCorrelationMatrix(Corr);
sam->generateSample(SamplerOption.Data());
}else{
TBasicSampling *sam = new TBasicSampling(tds,SamplingType.Data(),m);
sam->setCorrelationMatrix(Corr);
sam->generateCorrSample(SamplerOption.Data());
}
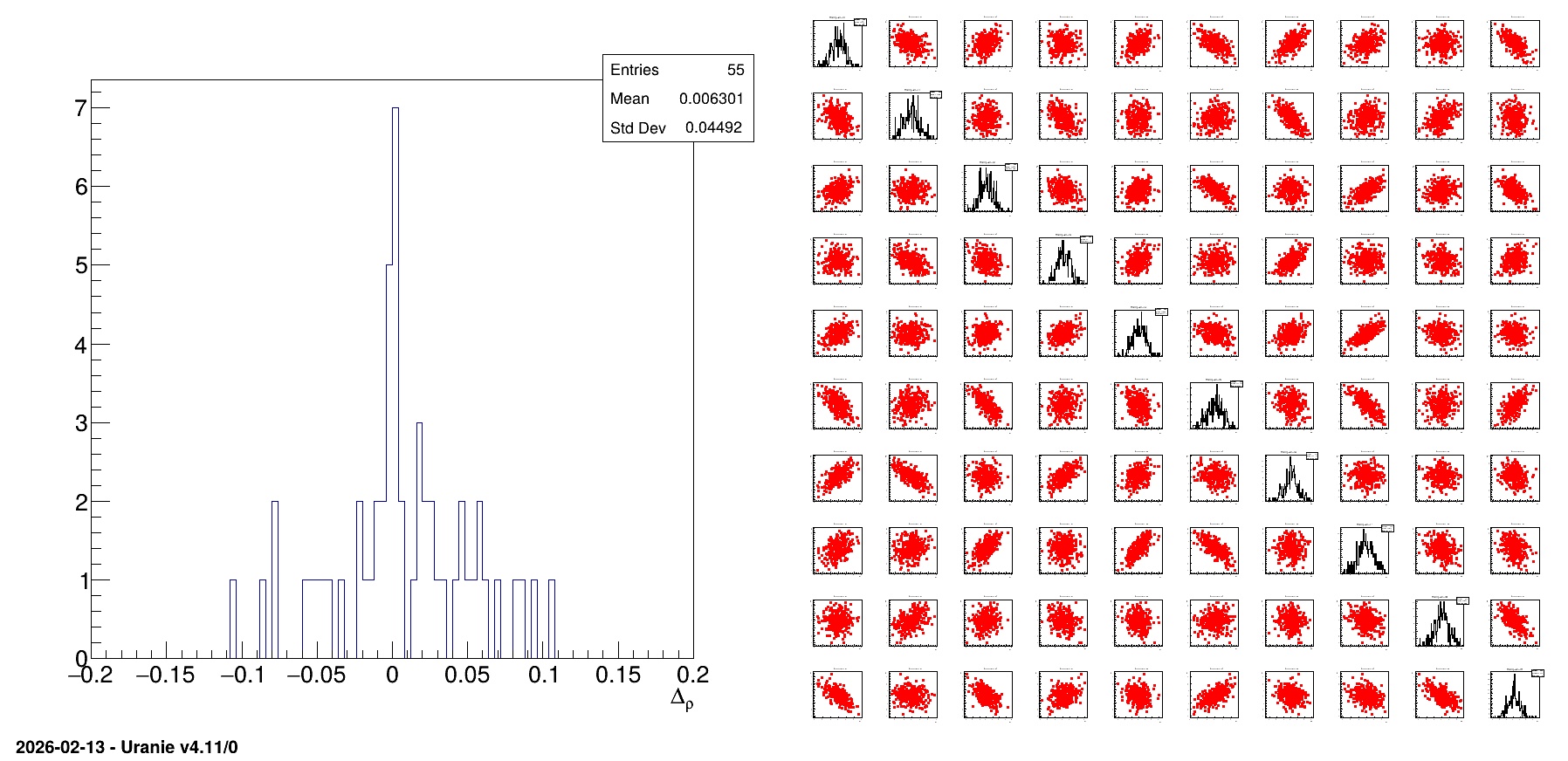
Finally the obtained design-of-experiments is shown along with the residual of all the correlation coefficients (difference between the target values and the obtained ones). This is shown in Figure 13.17 and can be used to compare the performance of the samplers.
13.3.14.3. Graph
Figure 13.17 Graph of the macro “samplingSingularCorrelationCase.C”