13.7.13. Macro “modelerbuildSimpleGPEstimWithCov.py”
13.7.13.1. Objective
This macro is the one described in Prediction of a new data set, global approach, to create and use a
simple gaussian process, whose training (utf_4D_train.dat) and testing (utf_4D_test.dat) database
can both be found in the document folder of the Uranie installation
(${URANIESYS}/share/uranie/docUMENTS). It uses the global approach, computing the input covariance
matrix and translating it to the prediction covariance matrix.
13.7.13.2. Macro Uranie
"""
Example of Gaussian Process building with estimation using full data (covariance)
"""
from URANIE import DataServer, Modeler
import ROOT
# Load observations
tdsObs = DataServer.TDataServer("tdsObs", "observations")
tdsObs.fileDataRead("utf_4D_train.dat")
# Construct the GPBuilder
gpb = Modeler.TGPBuilder(tdsObs, # observations data
"x1:x2:x3:x4", # list of input variables
"y", # output variable
"matern1/2") # name of the correlation function
# Search for the optimal hyper-parameters
gpb.findOptimalParameters("ML", # optimisation criterion
100, # screening design size
"neldermead", # optimisation algorithm
500) # max. number of optim iterations
# Construct the kriging model
krig = gpb.buildGP()
# Display model information
krig.printLog()
# Load the data to estimate
tdsEstim = DataServer.TDataServer("tdsEstim", "estimations")
tdsEstim.fileDataRead("utf_4D_test.dat")
# Reducing DB to 1000 first event (leading to cov matrix of a million value)
nS = 1000
tdsEstim.exportData("utf_4D_test_red.dat", "", "tdsEstim__n__iter__<="+str(nS))
# Reload reduced sample
tdsEstim.fileDataRead("utf_4D_test_red.dat", False, True)
krig.estimateWithCov(tdsEstim, # data to estimate
"x1:x2:x3:x4", # list of the input variables
"yEstim:vEstim", # name given to the model's outputs
"y", # name of the true reference
"DummyOption") # options
cTwo = None
# Produce residuals plots if true information provided
if tdsEstim.isAttribute("_Residuals_"):
cTwo = ROOT.TCanvas("c2", "c2", 1200, 800)
cTwo.Divide(2, 1)
cTwo.cd(1)
# Usual residual considering uncorrated input points
tdsEstim.Draw("_Residuals_")
cTwo.cd(2)
# Corrected residuals, taking full prediction covariance matrix
tdsEstim.Draw("_uncorrResiduals_")
# Retrieve all the prediction covariance coefficient
tdsEstim.getTuple().SetEstimate(nS*nS) # allocate the correct size
# Get a pointer to all values
tdsEstim.getTuple().Draw("_CovarianceMatrix_", "", "goff")
cov = tdsEstim.getTuple().GetV1()
# Put these in a matrix nicely created
Cov = ROOT.TMatrixD(nS, nS)
Cov.Use(0, nS-1, 0, nS-1, cov)
# Print it if size is reasonnable
if nS < 10:
Cov.Print()
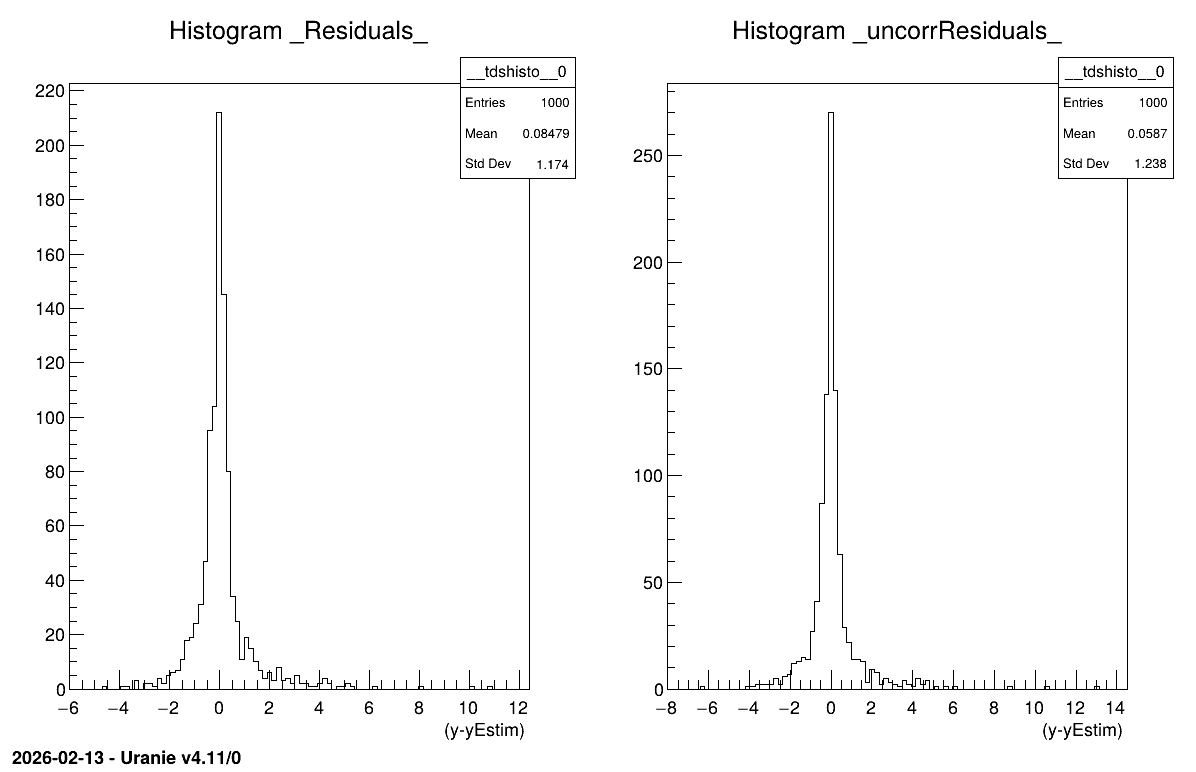
13.7.13.3. Graph
Figure 13.44 Graph of the macro “modelerbuildSimpleGPEstimWithCov.py”