13.11.1. Macro “calibrationMinimisationFlowrate1D.py”
13.11.1.1. Objective
The goal here is to calibrate the parameter \(H_l\) within the flowrate model, while varying only two
inputs (\(r_{\omega}\) and \(L\)). The remaining variables are fixed to the following values: \(r=25050\),
\(T_u=89335\), \(T_l=89.55\), \(H_u=1050\), \(K_{\omega}=10950\). The context of this example has already been
presented in Use-case for this chapter, including the model (implemented
here as a C++ function) and the initial lines defining the TDataServer objects. This macro demonstrates how
to apply a simple minimisation approach using a Relauncher-based architecture.
13.11.1.2. Macro Uranie
"""
Example of calibration using minimisation approach on flowrate 1D
"""
from URANIE import DataServer, Relauncher, Reoptimizer, Calibration
import ROOT
# Load the function flowrateCalib1D
ROOT.gROOT.LoadMacro("UserFunctions.C")
# Input reference file
ExpData = "Ex2DoE_n100_sd1.75.dat"
# define the reference
tdsRef = DataServer.TDataServer("tdsRef", "doe_exp_Re_Pr")
tdsRef.fileDataRead(ExpData)
# define the parameters
tdsPar = DataServer.TDataServer("tdsPar", "tdsPar")
tdsPar.addAttribute(DataServer.TAttribute("hl", 700.0, 760.0))
tdsPar.getAttribute("hl").setDefaultValue(728.0)
# Create the output attribute
out = DataServer.TAttribute("out")
# Create interface to assessors
Model = Relauncher.TCIntEval("flowrateCalib1D")
Model.addInput(tdsPar.getAttribute("hl"))
Model.addInput(tdsRef.getAttribute("rw"))
Model.addInput(tdsRef.getAttribute("l"))
Model.addOutput(out)
# Set the runner
runner = Relauncher.TSequentialRun(Model)
# Set the calibration object
cal = Calibration.TMinimisation(tdsPar, runner, 1)
cal.setDistance("LS", tdsRef, "rw:l", "Qexp")
solv = Reoptimizer.TNloptSubplexe()
cal.setOptimProperties(solv)
cal.estimateParameters()
# Draw the residuals
canRes = ROOT.TCanvas("CanRes", "CanRes", 1200, 800)
padRes = ROOT.TPad("padRes", "padRes", 0, 0.03, 1, 1)
padRes.Draw()
padRes.cd()
cal.drawResiduals("Residuals", "*", "", "nonewcanvas")
Apart from the initial lines described in the section Use-case for this chapter, the
key step is to define the starting point of the minimisation. This can be achieved either by calling the
setStartingPoint method of the TNlopt class, or by assigning a default value to the parameter attributes.
In this example, it is done as follows:
tdsPar.getAttribute("hl").setDefaultValue(728.0)
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
# Create interface to assessors
Model = Relauncher.TCIntEval("flowrateCalib1D")
Model.addInput(tdsPar.getAttribute("hl"))
Model.addInput(tdsRef.getAttribute("rw"))
Model.addInput(tdsRef.getAttribute("l"))
Model.addOutput(out)
# Set the runner
runner = Relauncher.TSequentialRun(Model)
Once this setup is complete, the calibration object (TMinimisation) is created. As explained in
Recommended distance and likelihood functions, construction method, the first step is
to define the distance function (here the least squares distance) using setDistance. This method
also specifies the TDataServer containing the reference data, the names of the reference inputs, and the reference
variable against which the model output is compared. Finally, the optimisation algorithm is defined by creating
an instance of TNloptSubplexe, and the parameters are then estimated.
# Set the calibration object
cal = Calibration.TMinimisation(tdsPar, runner, 1)
cal.setDistance("LS", tdsRef, "rw:l", "Qexp")
solv = Reoptimizer.TNloptSubplexe()
cal.setOptimProperties(solv)
cal.estimateParameters()
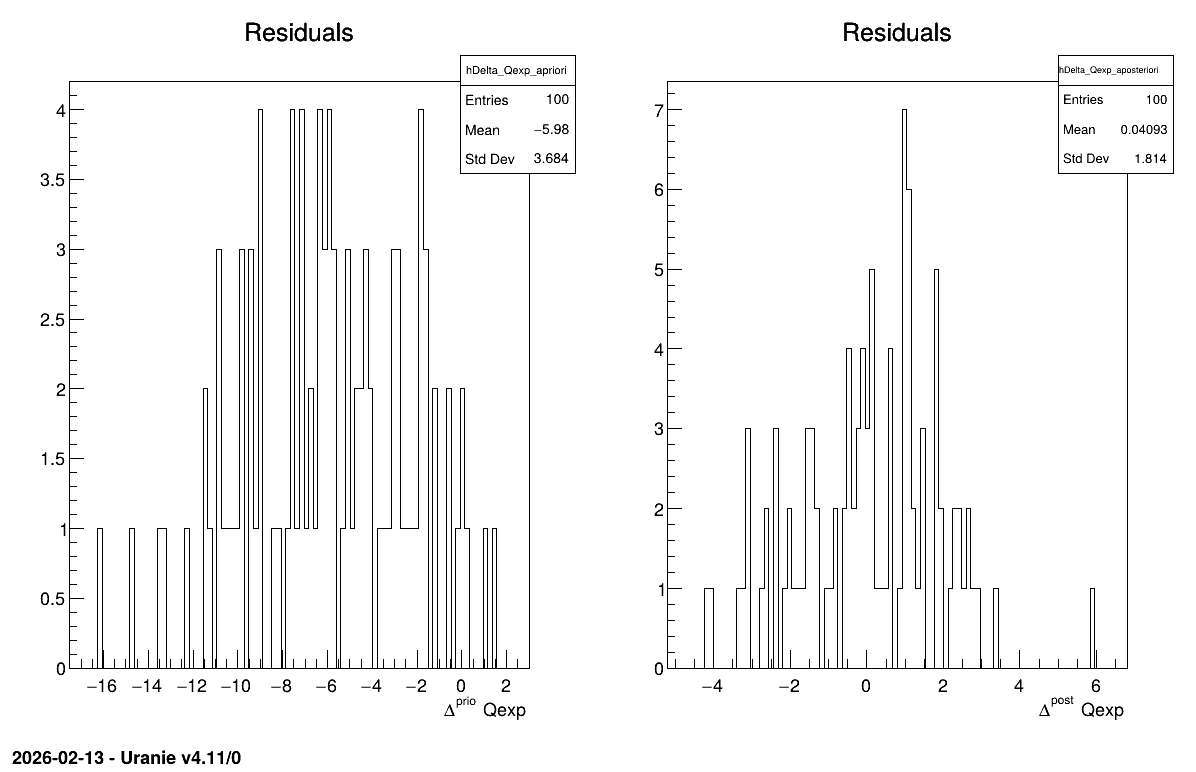
The final part demonstrates how to display the results. Since this method produces a point estimate, only a single value is obtained, which is always shown on the screen, as illustrated in Console. Another important aspect is to examine the residuals, as discussed in [Bla17]. This is illustrated in Figure 13.59, which shows the residuals of the a posteriori estimates, typically following a normal distribution.
# Draw the residuals
canRes = ROOT.TCanvas("CanRes", "CanRes", 1200, 800)
padRes = ROOT.TPad("padRes", "padRes", 0, 0.03, 1, 1)
padRes.Draw()
padRes.cd()
cal.drawResiduals("Residuals", "*", "", "nonewcanvas")
13.11.1.3. Console
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Thu Feb 12, 2026
|....:....|....:....|....:....|....:
.************************************************
* Row * tdsPar__n * hl.hl * agreement *
************************************************
* 0 * 0 * 749.72363 * 18.149368 *
************************************************
13.11.1.4. Graph
Figure 13.59 Residuals graph of the macro “calibrationMinimisationFlowrate1D.py”