Documentation
/ Manuel utilisateur en Python
: 
In a Bayesian framework, Markov Chain Monte Carlo (MCMC) methods are a powerful tool for calibration. They are especially valuable when the statistical model cannot be solved analytically, such as when the prior distribution has a complex structure or the model is nonlinear. Unlike many classical approaches, MCMC does not require the assumption of Gaussian errors: it remains applicable even when the likelihood is non-Gaussian.
Rather than providing a single "best-fit" solution (as in minimisation techniques), MCMC generates a collection of parameter samples that represent the full posterior distribution (similar to ABC methods). However, these methods also come at the cost of potentially high computational demand, since long sampling chains may be required to achieve convergence and reliable estimates. Users should therefore interpret results as distributions and ensure that convergence diagnostics are checked before drawing conclusions ( see [metho] for more details).
The usage of the TMCMC class can be summarised in a few key steps:
Prepare the data and the model:
The parameters to be calibrated must be instances of classes inheriting from
TStochasticAttribute;Select the assessor type and construct the
TMCMCobject with the appropriate likelihood function (see Section XI.6.1).
Set the algorithm properties:
Define optional behaviours;
Specify the uncertainty hypotheses via the dedicated methods (see Section XI.6.2).
Perform the estimate:
Run the estimate process;
Eventually continue it if the convergence is not reached (see Section XI.6.3).
Perform post-processing:
Investigate the quality of the samples through diagnostics and plots (see Section XI.6.4).
Analyse the results:
Extract the results and visualise them with the standard plotting tools (see Section XI.6.5).
The constructors available for creating an instance of the TMCMC class are detailed in
Section XI.2.3. As a reminder, the available prototypes are:
# Constructor with a runner
TMCMC(tds, runner, ns=1, option="")
# Constructor with a TCode
TMCMC(tds, code, ns=1, option="")
# Constructor with a function using Launcher
TMCMC(tds, fcn, varexpinput, varexpoutput, ns=1, option="")
Details about these constructors can be found in Section XI.2.3.1, Section XI.2.3.2, and Section XI.2.3.3
respectively for the TRun, TCode, and
TLauncherFunction-based constructor. In all cases, the number of iterations 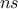 must be specified.
must be specified.
This class provides one specific option, which can be used to modify the default value of the a posteriori distribution returned by the algorithm. Two possible choices are available for obtaining the single-point estimate that best represents the distribution:
Mean of the distribution: this is the default option;
Mode of the distribution: the user must specify "mode" in the option field of the
TMCMCconstructor.
The default solution is straightforward, whereas the second requires internal smoothing of the distribution in order to obtain the best estimate of the mode.
In practice, the constructor, whichever one is chosen, initializes a folder named <foldername>MCMC_N</foldername>,
where N is an integer ensuring the folder name is available. This folder contains all the information related to
the MCMC calculation (the chain values, the algorithm used, the number of accepted samples, etc.), with each
chain stored in a separate file named MCMC_N_chain_M (for the M-th chain, starting from 0).
By default, only one chain is initialized, so the <foldername>MCMC_N</foldername> folder contains a single file
MCMC_N_chain_0. This folder will be duplicated once the setMultistart
method is called (see Section XI.6.2.4). The files are automatically loaded and saved
during the computation.
The next step is to construct the TDistanceLikelihoodFunction, a mandatory step that must
always immediately follow the constructor. For now, the only available likelihood function is the Gaussian, as it
is the most commonly used. It can be accessed through the classic setLikelihood
method using the function name "log-gauss", following the standard prototype (presented in
Section XI.2.2.1). As previously explained, in this case the weights provided to
the constructor correspond to the standard deviations of each observation. Advanced users also have the option
to define a custom likelihood by following the prototype:
setLikelihood(likelihoodFunc, tdsRef, input, reference, weight="")
Once the TMCMC instance is created along with its
TDistanceLikelihoodFunction, a few methods are available to tune the algorithms. All of
these methods are optional, as default values are provided. However, they should be applied in the following order:
Define the MCMC algorithm properties that are shared by all chains: MCMC algorithm, acceptation range, and information display interval (these can be set in any order).
Specify the number of chains to initialize.
Prepare the starting points and the initial standard deviations of the proposal distribution (these can be set in any order).
Several MCMC algorithms are available, and the desired algorithm can be selected using the setAlgo
method, which has the following prototype:
setAlgo(algoMCMC)
The method takes a single string argument that specifies the algorithm to use. Currently, the following options are supported:
Component-wise Metropolis-Hastings, specified with "MH_1D". This method updates one parameter at a time, which is useful when parameters have different influences on the model. By generating new candidates along on direction at a time, it prevents less influential parameters from being overshadowed by the more influential ones. This approach is generally slower than the classic Metropolis-Hastings;
Classic Metropolis-Hastings, specified with "MH_multiD" (default algorithm). This method generates candidates in all directions simultaneously, making it faster than the component-wise approach, but it may be less efficient when parameter influences vary widely.
Warning
As explained, this method should be done immediately after creating theTMCMC object
and assigning its likelihood.
As also explained in [metho], there are theoretical discussions on the acceptation rate expected, depending on the dimension of the parameter space for instance. As stated in some references (see [gelman1996efficient,roberts1997weak]) when well initialised, for a single-dimension problem, the acceptation rate could be around 44%, decreasing to about 23% as the dimensionality increases. By default, the exploration parameters are set during the initialisation step (see Section XI.6.2.5).
If one wants to change this, a possible approach is to use the setAcceptationRatioRange
method whose prototype is the following
setAcceptationRatioRange(lower, higher)
This prototype takes two arguments: the lower ( ) and higher (
) and higher (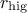 ) bounds. The idea is simply that after a certain number of estimate (called
batch, whose value is set to 100) the algorithm looks at the acceptation rate achieved
(actually this is computed for every configuration and kept in the final
) bounds. The idea is simply that after a certain number of estimate (called
batch, whose value is set to 100) the algorithm looks at the acceptation rate achieved
(actually this is computed for every configuration and kept in the final TDataServer object). If the lower and higher
bounds have been set, at the end of a batch, three cases are possible (when the acceptation rate is called
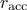 ):
):
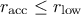 : the acceptation rate is too low compared to the desired range, this means the algorithm
is moving too far from the last accepted configuration. As a consequence, the variation range is reduced
by 10% to improve convergence.
: the acceptation rate is too low compared to the desired range, this means the algorithm
is moving too far from the last accepted configuration. As a consequence, the variation range is reduced
by 10% to improve convergence.
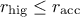 : the acceptation rate is too high compared to the desired range, this means that the
algorithm is not exploring the parameter space adequately, focusing only on a well-performing region.
As a consequence, the variation range is increased by 10% to encourage exploration.
: the acceptation rate is too high compared to the desired range, this means that the
algorithm is not exploring the parameter space adequately, focusing only on a well-performing region.
As a consequence, the variation range is increased by 10% to encourage exploration.
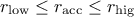 : the acceptance rate is within the desired range, no adjustment
is made.
: the acceptance rate is within the desired range, no adjustment
is made.
When the setAcceptationRatioRange method has been called, this process will be called
at the end of every batch. Otherwise, the default behaviour is to leave the
acceptation rate unchanged, as no boundaries have been set.
From the prototype and the behaviour discussed above, two rules must be followed when using this method:
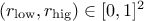 and
and 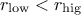 .
.
Finally, the evolution of the acceptance rate can be examined by plotting the relevant information, as discussed in Section XI.6.4.2.
The next method discussed here is fairly simple: a MCMC simulation may take a long time to complete, particularly when running code, regardless of the chosen architecture. This method changes the number of iterations between two output displays. The prototype is simply:
setNbDump(nbDump)
The method takes a single integer argument that specifies the interval (modulo) at which the algorithm displays its progress. With the default value of 1000, the output will look like this:
1000 events done
2000 events done
3000 events done
...
As already explained in [metho], initialising multiple chains can help assess the convergence of MCMC algorithms more reliably. Since these algorithms are inherently sequential (each state of the chain depending on the previous one), running several chains in parallel also makes it possible to exploit modern computational power more effectively. At present, this parallelization is not yet implemented (the chains are still computed sequentially) but it will be supported in a future release.
If you wish to initialise multiple chains, this should be done using the following prototype:
setMultistart(nb_multistart)
The method takes a single integer argument that specifies the number of chains to be initialised.
Warning
As explained, this method duplicates the file of the initial chain to prepare several chains, preserving the properties that have already been defined. For this reason, it should be performed immediately after selecting the MCMC algorithm, to ensure that all chains use the same algorithm. If the default algorithm is used, this step should be done after creating theTMCMC object and assigning its likelihood.
As already explained in [metho], the MCMC algorithm starts at a given point and then moves to a new configuration.
This means that two things must be specified for each parameter: a starting value, and a variation range used to
move to the next location. Since all the input parameters are stochastic (in other words, instances of classes
inheriting from TStochasticDistribution), the default
values are
randomly drawn for the starting point;
the theoretical standard deviation used as the variation range.
The default behaviour can easily be overridden by calling the setStartingPoints
and the setProposalStd methods with these prototypes:
# Prototype setStartingPoints
setStartingPoints(ichain, values)
# Prototype setProposalStd
setProposalStd(ichain, standDev)
The first argument of both prototypes is the chain index, an integer between -1 and  , where
, where 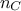 is the number of chains (indexed from 0 to
is the number of chains (indexed from 0 to  ). If -1 is chosen, all chains will be assigned the same
starting point (not recommended, since the purpose of using multiple chains is to initialise them at different
locations to verify convergence toward the same region) or the same standard deviation.
). If -1 is chosen, all chains will be assigned the same
starting point (not recommended, since the purpose of using multiple chains is to initialise them at different
locations to verify convergence toward the same region) or the same standard deviation.
The second argument of both prototypes is a vector containing either the initial values of the starting points (one
value for each parameter to calibrate) or the standard deviations (one for each parameter), which will be assigned to
the corresponding chain. This means that the vectors must have a size 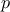 , corresponding to the number of parameters.
, corresponding to the number of parameters.
The computation can be performed using the standard estimateParameters method, which automatically
saves the chains in their associated files. Although it is generally unnecessary (since saving is done automatically), it
is still possible to manually export the results of a chain or reload them later using export_chain_MCMC
and read_chain_MCMC, whose prototypes are the following:
# Export one chain
export_chain_MCMC(fileName)
# Read one chain
read_chain_MCMC(fileName)
Both methods take a single string argument specifying the file name used for exporting or loading the chain results.
In most cases, these methods are not required. They become useful when a computation finishes without reaching
convergence and the user wishes to extend the run. This can be done with the continueCalculation
method, which executes additional iterations of the MCMC algorithm in order to try to achieve convergence. Its
prototype is:
continueCalculation(new_Ns)
This method takes a single integer argument that specifies the number of additional iterations to run. In this
case, it may be useful to reload one of the previously computed chains with
read_chain_MCMC before calling continueCalculation.
Regardless of which chain is loaded, the calculation will continue for all initialized chains.
Computing iterations does not guarantee that the chains have reached convergence and are properly sampling the posterior distribution. Although there is no exact way to prove convergence, several techniques can help assess the quality of the samples. Before plotting and analyzing the results, it is mandatory to check these diagnostics to determine whether the results are reliable or if the algorithm should be run for additional iterations (see Section XI.6.3). These techniques serve two main purposes:
Ensuring convergence of the chains: Convergence means that the chains have stabilized
around the same region. This must be checked by the user using the trace plot (see Section XI.6.4.1). It is also recommended to check the stability of the acceptance ratio, which should
typically lie between 20% and 50%, using the acceptance ratio plot (see Section XI.6.4.2).
Finally, the user must define the burn-in (also called warm-up), i.e., the number of
initial iterations discarded before the chain stabilizes. This is done with the setBurnin method:
setBurnin(burnin)
This method takes a single integer argument that specifies the number of iterations to remove (non-converged iterations).
When multiple chains are initialized (at least 4), it is also possible to compute the Gelman-Rubin statistic, which compares intra- and inter-chain variances (see Section XI.6.4.3). Values close to 1 indicate good convergence.
Ensuring approximate independence of posterior samples: Because Markov chains generate
dependent samples, there is a risk that successive samples are correlated and do not explore the posterior distribution
effectively. To assess this, the Effective Sample Size (ESS) can be computed with the
diagESS method (see Section XI.6.4.4), which provides the equivalent
number of independent samples. This method also suggests an appropriate lag value, i.e., the number of iterations to skip
before selecting the next uncorrelated sample. The lag is set using the setLag method:
setLag(lag)
The method takes a single integer argument that specifies the lag value.
Warning
Setting lag and burn-in values with thesetBurnin and setLag methods will
affect most drawing methods, and a line will indicate which default cut and lag were applied to produce the plot. The only
exception is the residuals plot, since the a posteriori residuals are computed during
estimateParameters, before any burn-in or lag is applied. To re-estimate residuals with a specific
lag or burn-in, use the estimateCustomResiduals method (see Section XI.2.3.5). To remove the current cut and lag, use the clearDefaultCut
method, which requires no arguments and clears both lag and burn-in:
clearDefaultCut()
This method is used to visualise the trace of the chains, i.e., the evolution of parameter values across iterations. It should be used to check the stability of the chains (stability of the mean and variance, see [metho]), and to detect potential autocorrelation (when the chain moves too slowly within the target distribution).
Thus, trace plots provide a first indication of the appropriate burn-in (the number of iterations to discard before stability is reached) and lag (the number of iterations to skip to reduce autocorrelation). The prototype is:
drawTrace(sTitre, variable = "*", option = "")
This method takes up to three arguments, two of which are optional:
- sTitre
the title of the plot (an empty string is allowed);
- variable (optional)
a list of parameter names to be drawn, separated by colons ":". The default "*" draws all parameters;
- option (optional)
a list of options, separated by commas "," to adjust the plotting behavior:
"nonewcanvas": draw on the current canvas (instead of creating a new one);
"vertical": if multiple parameters are plotted, display them stacked vertically (one per row). By default, plots are arranged horizontally, side by side.
Returning to the example in Section XIV.12.5, the trace plots (shown in Figure XIV.104) reveal that the very beginning of the chain is unstable, likely because initialization was far from the most probable value. Afterward, the behavior stabilizes, oscillating around what appears to be the most probable region. Based on this, a small burn-in of about 10 iterations could be chosen.
Looking at the acceptance ratio plot (see Figure XIV.105),
however, suggests that the burn-in should be slightly larger. Therefore, we set a value of 50 for the burn-in using
the setBurnin method (see Section XI.6.4). When a trace plot is
drawn after defining a burn-in with setBurnin, the burn-in region is indicated by a black
dotted line.
This method is used to visualise the evolution of the acceptation ratio across iterations. It should be used to check the stability of the chains, since an unstable acceptation ratio indicates that the chains may not have converged. It also allows the user to verify that the acceptation ratio stabilizes within the desired range (between 20% and 50%), which ensures good exploration of the target density while retaining a sufficient number of samples.
Thus, acceptation ratio plots provide useful guidance for determining an appropriate burn-in (the number of iterations to discard before stability is reached) and for selecting a suitable initial standard deviation of the proposal distribution to achieve an acceptation ratio within the desired range. The prototype is:
drawAcceptationRatio(sTitre, variable = "*", option = "")
This method takes up to three arguments, two of which are optional:
- sTitre
the title of the plot (an empty string is allowed);
- variable (optional)
a list of parameter names to be drawn, separated by colons ":". The default "*" draws all parameters;
- option (optional)
a list of options, separated by commas "," to adjust the plotting behavior:
"nonewcanvas": draw on the current canvas (instead of creating a new one);
"vertical": if multiple parameters are plotted, display them stacked vertically (one per row). By default, plots are arranged horizontally, side by side.
Returning to the example in Section XIV.12.5, the acceptation ratio plots (shown in
Figure XIV.105) reveal that
the acceptation ratio is unstable during the first iterations, likely because the chains were initialised far from the
most probable value. Afterward, the acceptation ratio stabilizes, oscillating around what appears to be an asymptote. Based
on this, a burn-in of about 50 can be considered appropriate. When a acceptation ratio plot is drawn after defining a
burn-in with setBurnin, the burn-in region is indicated by a black dotted line.
Once the trace and acceptance ratio plots have been analysed, and an initial burn-in value has been chosen, the Gelman-Rubin diagnostic can be used to verify the convergence of the chains. This diagnostic compares the intra- and inter-chain variances (see [metho] for more details).
The method diagGelmanRubin has the following prototype:
diagGelmanRubin()
This method takes no arguments, as it is called directly from the TCalibration object. For
example, in
Section XIV.12.5, the Gelman-Rubin
diagnostic is computed with:
GelmanRubin_values = cal.diagGelmanRubin()
GelmanRubin_values["hl"] # To extract the Gelman-Rubin statistic for parameter hl
The returned object GelmanRubin_values is an unordered map where each parameter name (e.g., "hl")
is associated with its Gelman-Rubin statistic value. The results are also printed in the console
(see Section XIV.12.1.3), accompanied by a short generic commentary interpreting the statistic.
In this example, the Gelman-Rubin diagnostic confirms good convergence of the chains (and thus the suitability of the chosen burn-in value). In other cases, where the statistics indicate poorer convergence, it may be necessary to run additional iterations (see Section XI.6.3) or to adjust the burn-in value (see Section XI.6.4).
Once convergence of the chains has been assessed using the trace plot (see Section XI.6.4.1), the acceptance ratio plot (see Section XI.6.4.2), and the Gelman-Rubin diagnostic (see Section XI.6.4.3), the next step is to evaluate the autocorrelation of the samples. If samples are highly correlated, the posterior distribution may not be properly explored (e.g., chains could remain trapped in a mode).
A common way to assess this is to compute the Effective Sample Size (ESS), which estimates how many samples can effectively be considered as independent among the available ones. ESS thus indirectly indicates the appropriate lag (the number of iterations to skip to obtain approximately uncorrelated samples). More details are provided in [metho].
The method diagESS has the following prototype:
diagESS()
This method takes no arguments, as it is called directly from the TCalibration object. For
example, in
Section XIV.12.5, the ESS diagnostic
is computed with:
ESS_values = cal.diagESS()
ESS_values["hl"][0] # To extract the ESS statistic for parameter hl computed on the first chain
The returned object ESS_values is an unordered map where each parameter name (e.g., "hl")
is associated with a vector of ESS values, one per chain. The results are also printed in the console (see Section XIV.12.1.3).
In this example, the ESS diagnostic confirms that each parameter has several hundred (at least 200) effectively uncorrelated
samples. It is therefore recommended to set the lag to 1 using setLag method (see Section XI.6.4), which indicates that the samples are sufficiently uncorrelated.
If the ESS is too small, it may be necessary to run additional iterations (see Section XI.6.3) or to adjust the initial standard deviation of the proposal distribution, which may be too small, causing the chain to move too slowly (see Section XI.6.2.5).
Once the convergence of the chains has been assessed and thinning applied, the results can be analysed in more detail. While trace plots are already informative results on their own, several additional methods are available. Among the implemented tools, it is possible to display pairwise parameter traces (see Section XI.6.5.1), visualise the posterior distributions (see Section XI.6.5.2), and examine the residual distribution (see Section XI.6.5.3). These methods are described in the following sections:
This method is used to visualise the trajectory of the chains across iterations by examining two parameters at a time. It can help verify that the chains uniformly explore the distribution and that they converge toward the same posterior. It also highlights potential covariance between pairs of parameters in the posterior distribution.
Although this method can be used as a diagnostic tool, it is less effective for assessing chain stability or detecting autocorrelation. For those purposes, standard trace plots remain preferable.
The prototype is:
draw2DTrace(sTitre, variable, option = "")
This method takes up to three arguments, two of which are optional:
- sTitre
the title of the plot (an empty string is allowed);
- variable (optional)
a list of exactly two parameter names to be drawn, separated by colons ":". For example,
"t0:t1"is a valid argument to plot the chains oft0andt1together on a 2D trace;- option (optional)
a list of options, separated by commas "," to adjust the plotting behavior:
"nonewcanvas": draw on the current canvas (instead of creating a new one);
Returning to the example in Section XIV.12.6, the 2D trace plots (shown in Figure XIV.110) clearly show the four chains converging toward the posterior distribution located in the center of the plot. The distribution appears to be uniformly covered by a sufficient number of samples, without any obvious trend suggesting covariance between the two parameters.
The posterior distributions of the parameters can be visualised using the dedicated instance of
drawParameters, whose prototype is identical to that discussed in Section XI.2.3.6.
void drawParameters(sTitre, variable = "*", option = "")
The only difference from the standard drawParameters method in
TCalibration is that this version automatically accounts for the burn-in period and the lag
(see Section XI.6.4 for details). Specifically, the first samples of each chain are
discarded according to the burn-in value, and the remaining samples are thinned using the specified lag (i.e., one
sample is retained out of every lag iterations).
An example of this function is shown in Figure XIV.112.
The residuals can be visualised with the standard drawResidues method of any
TCalibration object. Its arguments and options have already been detailed in
Section XI.2.3.6, and its prototype is recalled here for convenience:
drawResiduals(sTitre, variable = "*", select = "1>0", option = "")
Since no specific implementation has been added for MCMC calibration, the main point to clarify is how the a priori and a posteriori configurations are defined:
A priori configuration: obtained from the parameter initialisation. This can either be provided explicitly by the user through the method described in Section XI.6.2.5, or, if not specified, generated from a random draw according to the given A priori probability density.
A posteriori configuration: by default, the mean of the posterior distribution is used (as discussed in the constructor section of this class, see Section XI.6.1). If the option "mode" is specified when constructing the
TMCMCobject, then the posterior mode is estimated and used instead.
An example of this function is shown in Figure XIV.111.
Two examples are also provided in the use-case section (see Section XIV.12.5 and Section XIV.12.6).






