Documentation
/ Manuel utilisateur en C++
: 
This method is fairly simple from an algorithm point of view as it consists mainly of the analytical formulation of the posterior distribution under assumptions: the problem can be considered linear and the prior distributions are normally distributed (or non-informative/flat, as discussed in [metho]).
In practice, this technique is applied by following the procedure provided in Section XI.2 with one important difference, however: the code or function passed through the
constructor of the TLinearBayesian object is not strictly necessary. The parameter estimate
is analytical so the main point of providing an assessor is to get both the a
priori and a posteriori residuals distributions.
The usage of the TLinearBayesian class can be summarised in a few key steps:
Prepare the data and the model:
Select the assessor type to be used and construct the
TLinearBayesianobject with the appropriate likelihood function viasetLikelihoodmethod (see Section XI.4.1).
Set the algorithm properties:
Provide the input covariance matrix, i.e., the reference observation covariance (in [metho], this corresponds to
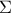 ). This
step is mandatory, as the covariance matrix is used to compute the posterior distribution, as discussed in
[metho];
). This
step is mandatory, as the covariance matrix is used to compute the posterior distribution, as discussed in
[metho];
Specify the name of the regressors. This is also a key step as a regressor can be an input variable, but also any function of one or several input variables. This is discussed in Section XI.4.2;
A transformation function may be provided, although this is optional. This is discussed in Section XI.4.3.1.
Perform the estimate and analyse the results:
Run the estimate process;
Extract the results and visualise them with the standard plotting tools (see Section XI.4.3).
The constructors available for creating an instance of the TLinearBayesian class are
those detailed in Section XI.2.3. As a reminder, the available prototypes are:
// Constructor with a runner
TLinearBayesian(TDataServer *tds, TRun *runner, Int_t ns=1, Option_t *option="");
// Constructor with a TCode
TLinearBayesian(TDataServer *tds, TCode *code, Int_t ns=1, Option_t *option="");
// Constructor with a function using Launcher
TLinearBayesian(TDataServer *tds, void (*fcn)(Double_t*,Double_t*), const char *varexpinput, const char *varexpoutput, int ns=1, Option_t *option="");
TLinearBayesian(TDataServer *tds, const char *fcn, const char *varexpinput, const char *varexpoutput, int ns=1, Option_t *option="");
Details about these constructors can be found in Section XI.2.3.1, Section XI.2.3.2, and Section XI.2.3.3
respectively for the TRun, TCode, and
TLauncherFunction-based constructor. In all cases, the number of samples 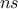 is set to 1 by default, and modifying it
has no effect on the results, since it returns the analytical distributions. This class does not define any
specific options.
is set to 1 by default, and modifying it
has no effect on the results, since it returns the analytical distributions. This class does not define any
specific options.
The final step is to construct the TDistanceLikelihoodFunction, a mandatory step that must
always immediately follow the constructor. This can be done via the setLikelihood method (following
the prototype presented in Section XI.2.2.1).
Warning
The Analytical Linear Bayesian Estimation method relies on the assumption that residuals are Gaussian (centered) with a defined covariance matrix (see Section XI.4.2). As a result, the likelihood is not a free choice but is determined by these assumptions. It is important to note that thesetLikelihood method is used only for residual calculation,
to compare the prior and posterior. Regardless of the initial choice of likelihood, the function is locally redefined
so that the computation is performed via the lin_gauss function using matrix multiplication (through the
TMahalanobisDistance method).
Although not recommended, it is still possible to define a custom likelihood function:
void setLikelihood(TDistanceLikelihoodFunction *likelihoodFunc, TDataServer *tdsRef, const char *input, const char *reference, const char *weight="");
Once the TLinearBayesian instance is created along with its
TDistanceLikelihoodFunction, two methods must be called before performing
parameter estimate. These methods are mandatory, as they define the analytical formula used
to obtain the Gaussian parameter values of the a posteriori distribution
(see [metho]).
The first method (although the order is not important) is setRegressor,
whose prototype is
void setRegressor(const char *regressorname);
The only argument is regressorname, a string containing the list of regressor names separated by
":". The method then performs two checks: it verifies that the number of regressors matches the number of parameters
to be calibrated and it checks that every regressor name provided matches one existing attribute in the reference
TDataServer (tdsref). If the observation TDataServer does not contain the regressors (when the input file is loaded)
these attributes must be constructed from scratch, either with TAttributeFormula
or by using another dedicated assessor (as done in the use-case shown in Section XIV.11.3).
The other method is setObservationCovarianceMatrix whose prototype is
void setObservationCovarianceMatrix(TMatrixD &mat);
The only argument here is a TMatrixD whose content is the covariance matrix of the reference
observation data. Once again, this method will check two things:
the provided matrix must have the correct dimensions (both rows and columns must be equal to
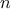 );
);
the provided matrix should be symmetrical;
Once these conditions are satisfied, estimate can proceed. One can find an example of how to use these methods in the use-case dedicated subsection (see Section XIV.11.3).
Finally, once the computation is complete there are three different kinds of results, with several ways to interpret and transform them. This section describes some important and specific aspects of these results.
The idea is that when you want to consider your model as linear, you may need to slightly transform it to ensure
proper linear behaviour and to express and compute the needed regressors. For this particular situation, the use-case
described in Section XIV.11.3 will be used. In this case, the flowrate function
should be linearised as follows:
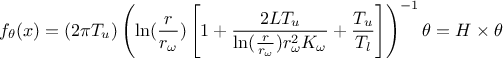
where the regressor can be expressed as
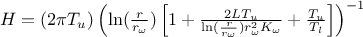 . From this, it is clear that we will be calibrating a newly
defined parameter
. From this, it is clear that we will be calibrating a newly
defined parameter 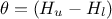 . Therefore, at some point, we will need to transform it back into
our parameter of interest.
. Therefore, at some point, we will need to transform it back into
our parameter of interest.
This is the reason why the setParameterTransformationFunction method has been implemented:
to transform the estimated parameters given the linear regressor, the observation covariance matrix, and the prior
distribution. Since the transformations, if they exist (they are optional), are expected to be carried out using
simple operations with constant values, they should affect only the mean vector and not the covariance matrix of
the posterior multivariate normal distribution. The prototype of this function is as follows:
void setParameterTransformationFunction(void (*fTransfoParam)(double *in, double *out));
Its only argument is a pointer to the transformation function. This function, used to obtain the transformed parameter values, takes two arguments: the input parameters, which are the raw values estimated from the analytical formula detailed in [metho], and the output parameters, which correspond to the desired transformed values. Both arguments are double arrays of length equal to the number of parameters.
The example provided in the use-case Section XIV.11.3 is simple as there is only one parameter to be estimated, which implies that both arguments are one-dimensional double arrays which should look like this:
void transf(double *x, double *res)
{
res[0] = 1050 - x[0]; // simply H_l = \theta - H_u
}
Once the estimate is done (via the classic estimateParameters method), the results can
be accessed by calling three methods, detailed below. All three functions share the same prototype: they take no
arguments and return a TMatrixD instance containing the corresponding information. The functions
are:
- getParameterValueMatrix
it returns a
TMatrixDwith one raw filled with the calibrated values of the parameters calculated from the analytical formula (one-dimensional);- getParameterCovarianceMatrix
it returns a
TMatrixDfilled with the covariance matrix of the estimated parameters (symmetric and (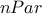 ,
,
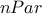 )-dimensional);
)-dimensional);
- getTransfParameterValueMatrix
it returns a
TMatrixDwith one raw filled with the transformed values of the parameters, in casesetParameterTransformationFunctionhas been correctly applied (one-dimensional).
Parameters can be plotted using a newly defined drawParameters instance, which shares the
same prototype as the original method described in Section XI.2.3.6.
void drawParameters(TString sTitre, const char *variable = "*", Option_t * option = "");
It takes up to three arguments, two of which are optional:
- sTitre
the title of the plot (an empty string is allowed);
- variable (optional)
a list of parameter names to be drawn, separated by colons ":". The default "*" draws all parameters;
- option (optional)
a list of options, separated by commas "," to adjust the plotting behavior:
"nonewcanvas": draw on the current canvas (instead of creating a new one);
"vertical": if multiple parameters are plotted, display them stacked vertically (one per row). By default, plots are arranged horizontally, side by side.
"apriori/aposteriori": draw only the a priori residuals or only the a posteriori residuals. If neither option is specified, both are displayed;
"transformed": this option specifies that the transformed values should be used as the mean vector of the multivariate normal posterior distribution.
The main difference compared to the standard drawParameters method in
TCalibration is that the plotted object are analytical functions.
In addition to the parameters, the residuals can be plotted using the standard drawResiduals
method, which remains unchanged (see Section XI.2.3.7).
computePredictionVariance method:
void computePredictionVariance(URANIE::DataServer::TDataServer *tdsPred, string outname);
- tdsPred
a
TDataServercontaining the new locations to be estimated, in which all regressors must be available in order to be able to compute the covariance matrix;- outname
the name of the attribute to be created, which will be filled with the diagonal elements (the variances) of the
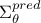 matrix.
matrix.






