Documentation
/ Manuel utilisateur en C++
: 
This section introduces a few examples of calibration in order to illustrate the different techniques introduced in Chapter XI, along with some of the available options.
The goal here is to calibrate the parameter 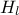 within the
within the flowrate model, while varying only two inputs
(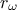 and
and
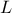 ). The remaining variables
are fixed to the following values:
). The remaining variables
are fixed to the following values: 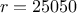 ,
, 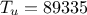 ,
, 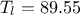 ,
, 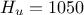 ,
, 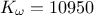 . The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the
. The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the TDataServer objects. This macro demonstrates how to apply a simple minimisation approach using
a Relauncher-based architecture.
{
// Load the function flowrateCalib1D
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="Ex2DoE_n100_sd1.75.dat";
// define the reference
TDataServer *tdsRef = new TDataServer("tdsRef","doe_exp_Re_Pr");
tdsRef->fileDataRead(ExpData.Data());
// define the parameters
TDataServer *tdsPar = new TDataServer("tdsPar","pouet");
tdsPar->addAttribute( new TAttribute("hl", 700.0, 760.0) );
tdsPar->getAttribute("hl")->setDefaultValue(728.0);
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib1D");
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
// Set the calibration object
TMinimisation *cal = new TMinimisation(tdsPar,runner,1);
cal->setDistance("LS",tdsRef,"rw:l","Qexp");
TNloptSubplexe solv;
cal->setOptimProperties(&solv);
cal->estimateParameters();
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *apad = new TPad("apad","apad",0, 0.03, 1, 1); apad->Draw(); apad->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
}
Apart from the initial lines described in the section Section XI.2.4, the key step
is to define the starting point of the minimisation. This can be achieved either by calling the
setStartingPoint method of the TNlopt class, or by assigning a
default value to the parameter attributes. In this example, it is done as follows:
tdsPar->getAttribute("hl")->setDefaultValue(728.0);
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib1D");
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
Once this setup is complete, the calibration object (TMinimisation) is created. As explained in Section XI.2.2.1, the first step is to define the distance function (here the least squares
distance) using setDistance. This method also specifies the TDataServer
containing the reference data, the names of the reference inputs, and the reference variable against which the model
output is compared. Finally, the optimisation algorithm is defined by creating an instance of
TNloptSubplexe, and the parameters are then estimated.
// Set the calibration object
TMinimisation *cal = new TMinimisation(tdsPar,runner,1);
cal->setDistance("LS",tdsRef,"rw:l","Qexp");
TNloptSubplexe solv;
cal->setOptimProperties(&solv);
cal->estimateParameters();
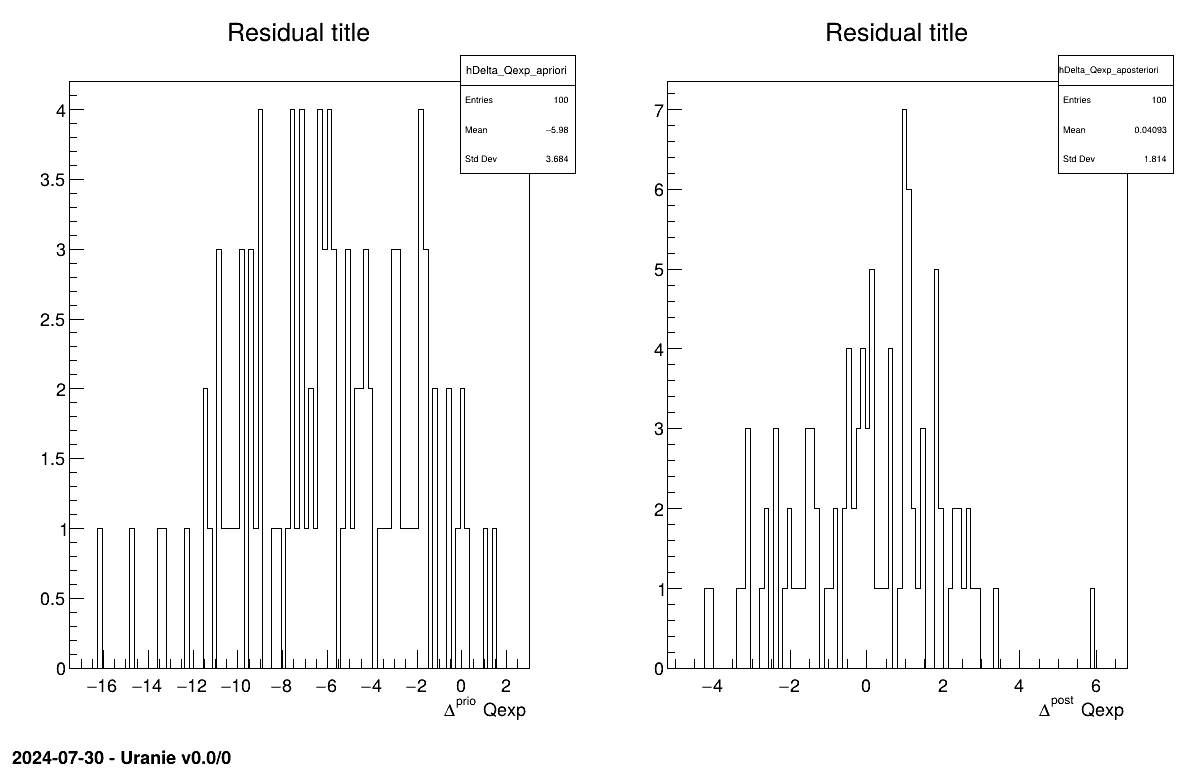
The final part demonstrates how to display the results. Since this method produces a point estimate, only a single value is obtained, which is always shown on the screen, as illustrated in Section XIV.11.1.3. Another important aspect is to examine the residuals, as discussed in [metho]. This is illustrated in Figure XIV.97, which shows the residuals of the a posteriori estimates, typically following a normal distribution.
// Draw the residuals
apad->Draw();
apad->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
Processing calibrationMinimisationFlowrate1D.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
|....:....|....:....|....:....|....:
.************************************************
* Row * tdsPar__n * hl.hl * agreement *
************************************************
* 0 * 0 * 749.72363 * 18.149368 *
************************************************
The goal here is to calibrate the parameter parameters 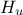 and
and 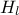 within the
within the flowrateCalib2D model (a two-dimensional version of the
flowrate model), while varying only two inputs
(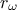 and
and
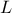 ). The remaining variables
are fixed to the following values:
). The remaining variables
are fixed to the following values: 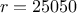 ,
, 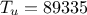 ,
, 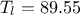 ,
, 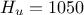 ,
, 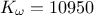 . The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the
. The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the TDataServer objects.
In addition to what has been presented in Section XIV.11.1, this macro introduces two new elements:
the use of Vizir instead of a simpler
TNloptSolver-inheriting instance;the discussion of problem identifiability, as introduced in [metho].
{
// Load the function flowrateCalib2DVizir
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="Ex2DoE_n100_sd1.75.dat";
// define the reference
TDataServer *tdsRef = new TDataServer("tdsRef","doe_exp_Re_Pr");
tdsRef->fileDataRead(ExpData.Data());
// define the parameters
TDataServer *tdsPar = new TDataServer("tdsPar","pouet");
tdsPar->addAttribute( new TAttribute("hu", 1020.0, 1080.0) );
tdsPar->addAttribute( new TAttribute("hl", 720.0, 780.0) );
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib2D");
Model->addInput(tdsPar->getAttribute("hu"));
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
// Set the calibration object
TMinimisation *cal = new TMinimisation(tdsPar,runner,1);
cal->setDistance("relativeLS",tdsRef,"rw:l","Qexp");
// Set optimisaiton properties
URANIE::Reoptimizer::TVizirGenetic solv;
solv.setSize(24,15000,100);
cal->setOptimProperties(&solv);
// ((URANIE::Reoptimizer::TVizir2*)cal->getOptimMaster()->setTolerance(1e-6);
cal->estimateParameters();
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1); padRes->Draw(); padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
// Draw the box plot of parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
tdsPar->getTuple()->SetMarkerStyle(20); tdsPar->getTuple()->SetMarkerSize(0.8);
tdsPar->Draw("hu:hl");
// Look at the correlation and statistic
tdsPar->computeStatistic("hu:hl");
TMatrixD corr=tdsPar->computeCorrelationMatrix("hu:hl");
corr.Print();
cout<<"hl is "<<tdsPar->getAttribute("hl")->getMean()<<" +- "<<tdsPar->getAttribute("hl")->getStd()<<endl;
cout<<"hu is "<<tdsPar->getAttribute("hu")->getMean()<<" +- "<<tdsPar->getAttribute("hu")->getStd()<<endl;
}
Much of this code has already been covered in the previous section Section XIV.11.1
(up to the sequential run). The main difference here is that the input parameter is now defined as a
TAttribute with boundaries specifying the space in which the algorithm will search.
tdsPar->addAttribute( new TAttribute("hu", 1020.0, 1080.0) );
tdsPar->addAttribute( new TAttribute("hl", 720.0, 780.0) );
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib2D");
Model->addInput(tdsPar->getAttribute("hu"));
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
Once this setup is complete, the calibration object (TMinimisation) is created. As explained
in Section XI.2.2.1, the first step is to define the distance function (here the
relative least squares distance) using setDistance. This method also specifies the TDataServer
containing the reference data, the names of the reference inputs, and the reference variable against which the model
output is compared. Finally, the optimisation algorithm is defined by creating an instance of
TVizirGenetic, and the parameters are then estimated.
// Set the calibration object
TMinimisation *cal = new TMinimisation(tdsPar,runner,1);
cal->setDistance("relativeLS",tdsRef,"rw:l","Qexp");
// Set optimisation properties
URANIE::Reoptimizer::TVizirGenetic solv;
solv.setSize(24,15000,100);
cal->setOptimProperties(&solv);
// ((URANIE::Reoptimizer::TVizir2*)cal->getOptimMaster()->setTolerance(1e-6);
cal->estimateParameters();
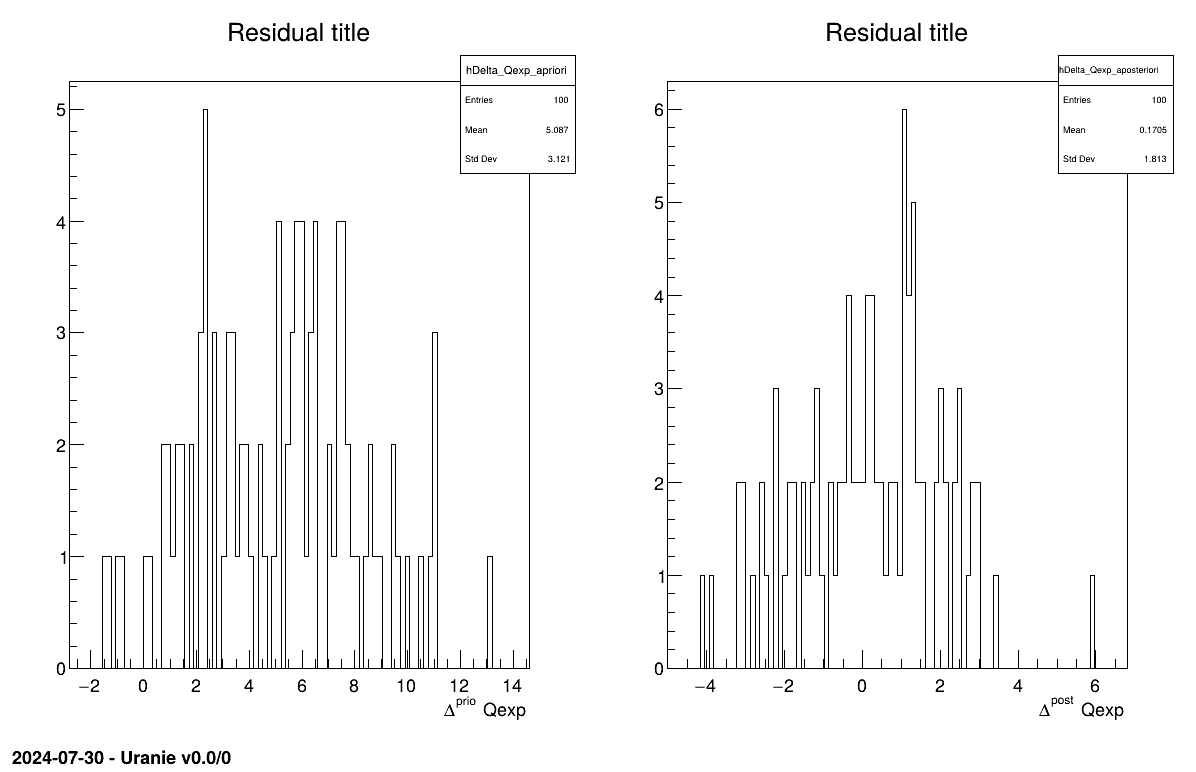
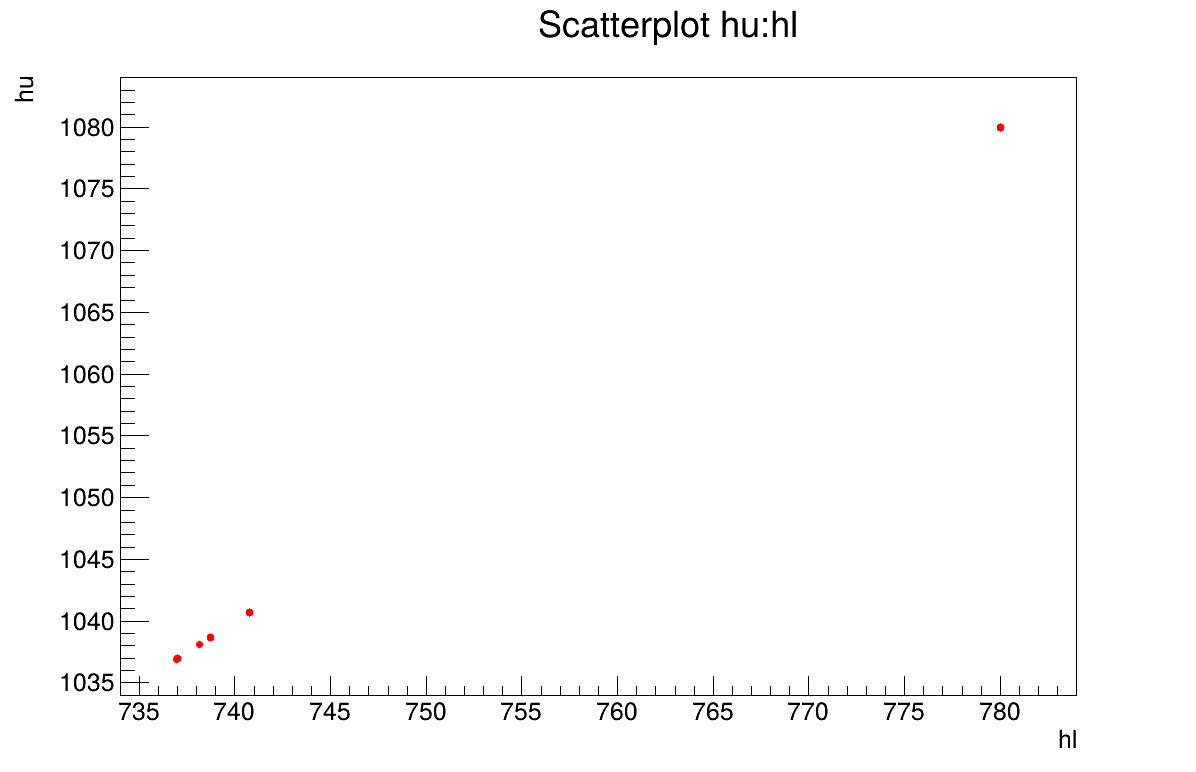
The final part demonstrates how to display the results. Since this method produces a point estimate, only a single value is obtained, which is always shown on the screen, as illustrated in Section XIV.11.2.3. Another important aspect is to examine the residuals, as discussed in [metho]. This is illustrated in Figure XIV.98, which shows the residuals of the a posteriori estimates, typically following a normal distribution. Finally, the parameter graph (Figure XIV.99) reveals a wide variety of possible solutions. This highlights a problem of identifiability, since infinitely many parameter combinations can lead to the same results, as further confirmed by the correlation matrix shown in Section XIV.11.2.3.
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
// Draw the box plot of parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
tdsPar->getTuple()->SetMarkerStyle(20);
tdsPar->getTuple()->SetMarkerSize(0.8);
tdsPar->Draw("hu:hl");
// Look at the correlation and statistic
tdsPar->computeStatistic("hu:hl");
TMatrixD corr=tdsPar->computeCorrelationMatrix("hu:hl");
corr.Print();
cout<<"hl is "<<tdsPar->getAttribute("hl")->getMean()<<" +- "<<tdsPar->getAttribute("hl")->getStd()<<endl;
cout<<"hu is "<<tdsPar->getAttribute("hu")->getMean()<<" +- "<<tdsPar->getAttribute("hu")->getStd()<<endl;
3
Processing calibrationMinimisationFlowrate2DVizir.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
first 100
Genetic 1
Generation : 1, rang max 23
Nb d'evaluation : 100, taille de la Z.P. : 0
Generation : 2, rang max 23
Nb d'evaluation : 465, taille de la Z.P. : 1
Generation : 3, rang max 8
Nb d'evaluation : 963, taille de la Z.P. : 6
Generation : 4, rang max 0
Nb d'evaluation : 1617, taille de la Z.P. : 24
Genetic converge 1617
************************************************************************************
* Row * tdsPar__n * hu.hu * hl.hl * agreement * rgpareto. * generatio *
************************************************************************************
* 0 * 0 * 1038.1302 * 738.13383 * 0.3639076 * 0 * 3 *
* 1 * 1 * 1080 * 780 * 0.3639083 * 0 * 3 *
* 2 * 2 * 1040.7058 * 740.77116 * 0.3639018 * 0 * 3 *
* 3 * 3 * 1079.9545 * 780 * 0.3639024 * 0 * 3 *
* 4 * 4 * 1040.7058 * 740.77116 * 0.3639018 * 0 * 1 *
* 5 * 5 * 1038.6638 * 738.70819 * 0.3639025 * 0 * 3 *
* 6 * 6 * 1040.7058 * 740.77116 * 0.3639018 * 0 * 3 *
* 7 * 7 * 1036.9776 * 736.98122 * 0.3639076 * 0 * 3 *
* 8 * 8 * 1036.9776 * 736.98122 * 0.3639076 * 0 * 3 *
* 9 * 9 * 1038.6638 * 738.70819 * 0.3639025 * 0 * 3 *
* 10 * 10 * 1036.9776 * 736.98122 * 0.3639076 * 0 * 2 *
* 11 * 11 * 1080 * 780 * 0.3639083 * 0 * 3 *
* 12 * 12 * 1036.9776 * 736.98122 * 0.3639076 * 0 * 3 *
* 13 * 13 * 1040.7058 * 740.77116 * 0.3639018 * 0 * 2 *
* 14 * 14 * 1080 * 780 * 0.3639083 * 0 * 3 *
* 15 * 15 * 1080 * 780 * 0.3639083 * 0 * 3 *
* 16 * 16 * 1038.6638 * 738.70819 * 0.3639025 * 0 * 3 *
* 17 * 17 * 1040.7058 * 740.77116 * 0.3639018 * 0 * 3 *
* 18 * 18 * 1080 * 780 * 0.3639083 * 0 * 3 *
* 19 * 19 * 1036.9776 * 736.98122 * 0.3639076 * 0 * 3 *
* 20 * 20 * 1036.9042 * 736.96108 * 0.3639019 * 0 * 3 *
* 21 * 21 * 1038.6638 * 738.70819 * 0.3639025 * 0 * 3 *
* 22 * 22 * 1080 * 780 * 0.3639083 * 0 * 3 *
* 23 * 23 * 1080 * 780 * 0.3639083 * 0 * 3 *
************************************************************************************
2x2 matrix is as follows
| 0 | 1 |
-------------------------------
0 | 1 1
1 | 1 1
hl is 752.445 +- 19.9463
hu is 1052.42 +- 19.9598
The goal here is to calibrate the parameter 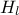 within the
within the flowrate model, while varying only two inputs
(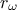 and
and
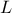 ). The remaining variables
are fixed to the following values:
). The remaining variables
are fixed to the following values: 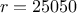 ,
, 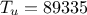 ,
, 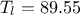 ,
, 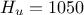 ,
, 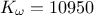 . The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the
. The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the TDataServer objects. This macro demonstrates how to apply a linear Bayesian estimation technique
using a Relauncher-based architecture.
{
// Load the function flowrateCalib1D
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="Ex2DoE_n100_sd1.75.dat";
// define the reference
TDataServer *tdsRef = new TDataServer("tdsRef","doe_exp_Re_Pr");
tdsRef->fileDataRead(ExpData.Data());
// define the parameters
TDataServer *tdsPar = new TDataServer("tdsPar","pouet");
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *Reg = new TCIntEval("flowrateModelnoH");
Reg->addInput(tdsRef->getAttribute("rw"));
Reg->addInput(tdsRef->getAttribute("l"));
Reg->addOutput(new TAttribute("H") );
TSequentialRun *runnoH = new TSequentialRun(Reg);
runnoH->startSlave();
if(runnoH->onMaster())
{
TLauncher2 l(tdsRef, runnoH);
l.solverLoop();
runnoH->stopSlave();
}
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib1D");
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
// Set the covariance matrix of the input reference
double sd=tdsRef->getValue("sd_eps",0);
TMatrixD mat(100,100);
for(unsigned int ival=0; ival<tdsRef->getNPatterns(); ival++)
mat(ival,ival)=(sd*sd);
// Set the calibration object
TLinearBayesian *cal = new TLinearBayesian(tdsPar,runner,1,"");
cal->setLikelihood("gauss_lin",tdsRef,"rw:l","Qexp");
cal->setObservationCovarianceMatrix(mat);
cal->setRegressorName("H");
cal->setParameterTransformationFunction(transf);
cal->estimateParameters();
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1); padRes->Draw(); padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
// Draw the parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1); padPar->Draw(); padPar->cd();
cal->drawParameters("Parameters","*","nonewcanvas,transformed");
}
Much of this code has already been covered in the previous section Section XIV.11.1.2
(up to the sequential run). The main difference here is that the input parameter is now defined as a
TStochasticDistribution, representing the a priori chosen distribution.
In this example, it can be either a TNormalDistribution or a TUniformDistribution
(see [metho]), with the latter being selected here:
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
Another difference from the previous example is that the method must be slightly adapted to obtain values for the
regressor. As discussed in [metho], linear Bayesian estimation can be applied when the model is approximately linear.
This requires linearising the flowrate function, as demonstrated here:
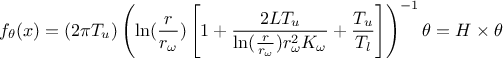
Here, the regressor can be expressed as 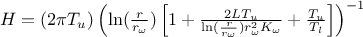 .
From this expression, it is clear that we will be calibrating a newly defined parameter
.
From this expression, it is clear that we will be calibrating a newly defined parameter 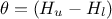 , which must later be converted back into the parameter of interest. To obtain the regressor
estimation, we simply use another C++ function,
, which must later be converted back into the parameter of interest. To obtain the regressor
estimation, we simply use another C++ function, flowrateModelnoH together with the standard
Relauncher approach:
// Create interface to assessors
TCIntEval *Reg = new TCIntEval("flowrateModelnoH");
Reg->addInput(tdsRef->getAttribute("rw"));
Reg->addInput(tdsRef->getAttribute("l"));
Reg->addOutput(new TAttribute("H") );
TSequentialRun *runnoH = new TSequentialRun(Reg);
runnoH->startSlave();
if(runnoH->onMaster())
{
TLauncher2 launch(tdsRef, runnoH);
launch.solverLoop();
runnoH->stopSlave();
}
This method also requires the input covariance matrix. The provided dataset
(Ex2DoE_n100_sd1.75.dat) contains an estimate of the uncertainty affecting the reference
measurements. Since this uncertainty is constant across all samples, the input covariance matrix is diagonal,
with each diagonal element equal to the square of the standard deviation, as illustrated below:
// Set the covariance matrix of the input reference
double sd=tdsRef->getValue("sd_eps",0);
TMatrixD mat(100,100);
for(unsigned int ival=0; ival<tdsRef->getNPatterns(); ival++)
mat(ival,ival)=(sd*sd);
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
The calibration object is then created using a Mahalanobis distance function, which is used here for illustrative purposes (see Section XI.4 for more details). The three key steps are:
Providing the input covariance matrix via the
setObservationCovarianceMatrixmethod;Specifying the regressorsâ names using
setRegressorName;Defining the parameter transformation function (optional) with
setParameterTransformationFunction.
The final step is more involved: since we are calibrating 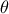 , we need to recover the corresponding parameter value at the end.
This is achieved by providing a C++ function that transforms the parameter estimated from the linearisation back into
the parameter of interest. For illustration, this is implemented in
, we need to recover the corresponding parameter value at the end.
This is achieved by providing a C++ function that transforms the parameter estimated from the linearisation back into
the parameter of interest. For illustration, this is implemented in UserFunctions.C via the
transf function, shown below:
void transf(double *x, double *res)
{
res[0] = 1050 - x[0]; // simply H_l = \theta - H_u
}
The complete block of code discussed in this section is as follows:
// Set the calibration object
TLinearBayesian *cal = new TLinearBayesian(tdsPar,runner,1,"");
cal->setLikelihood("Mahalanobis",tdsRef,"rw:l","Qexp");
cal->setObservationCovarianceMatrix(mat);
cal->setRegressorName("H");
cal->setParameterTransformationFunction(transf);
cal->estimateParameters();
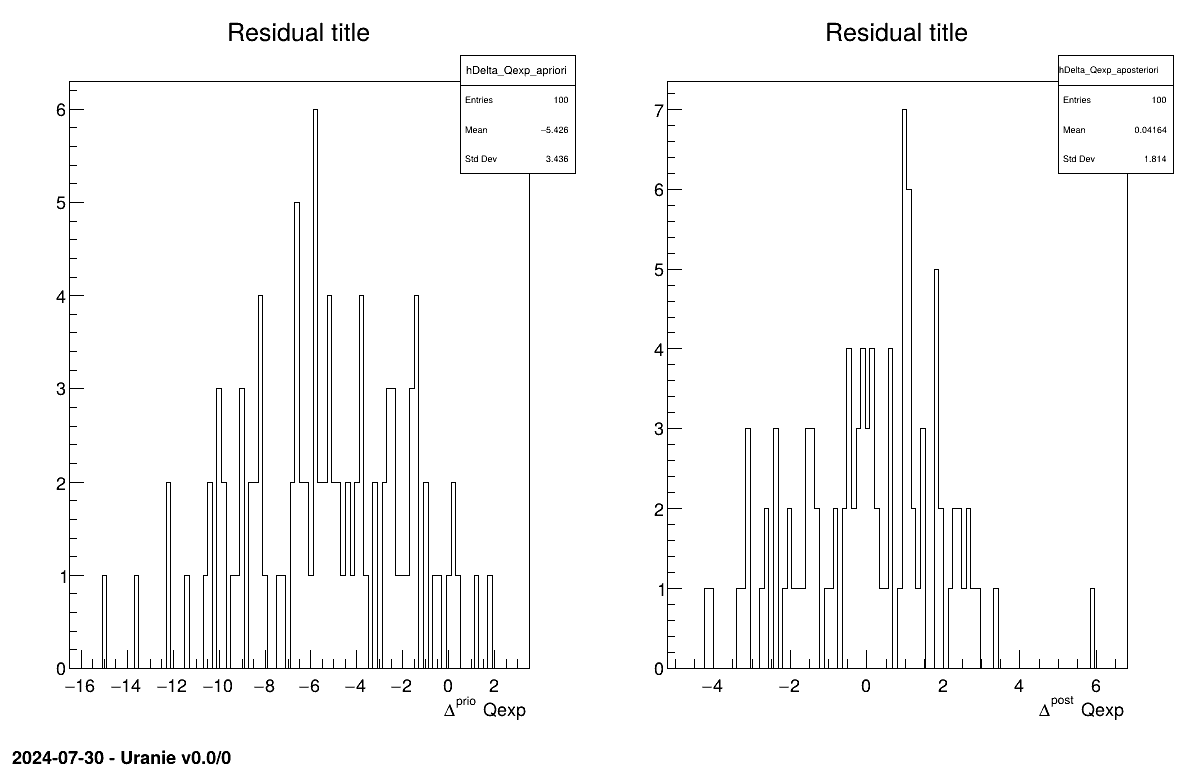
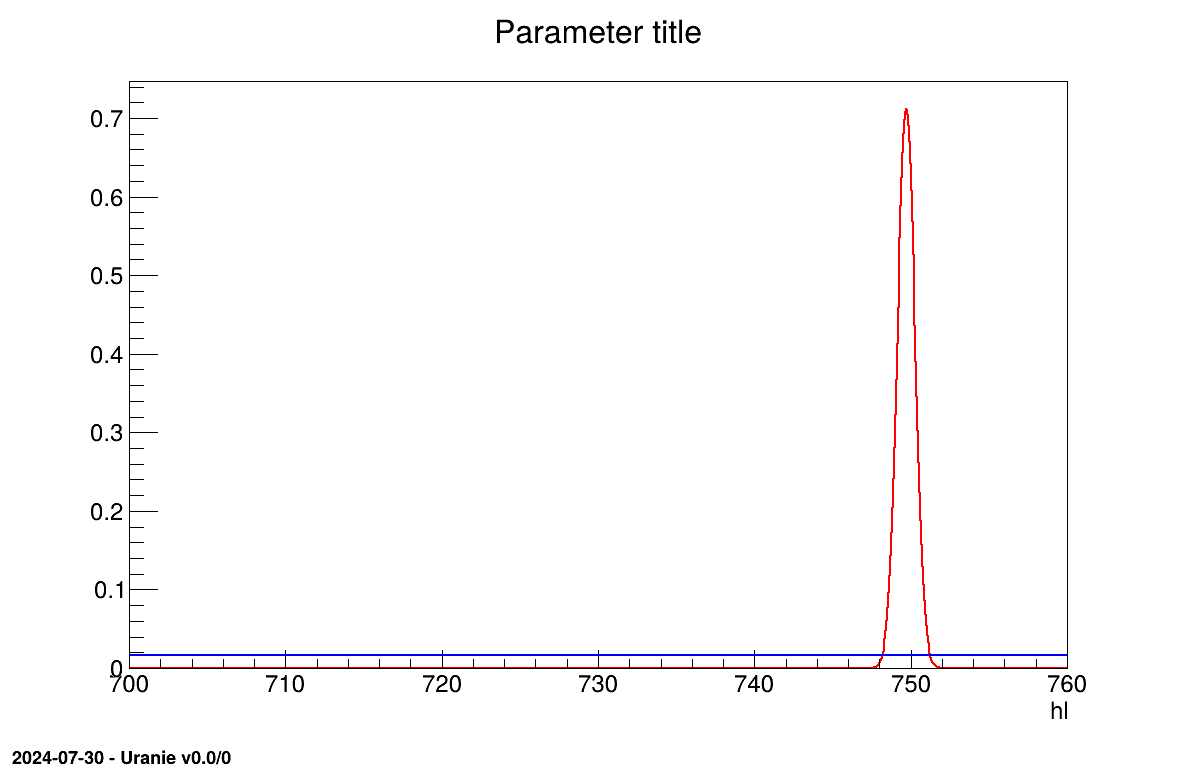
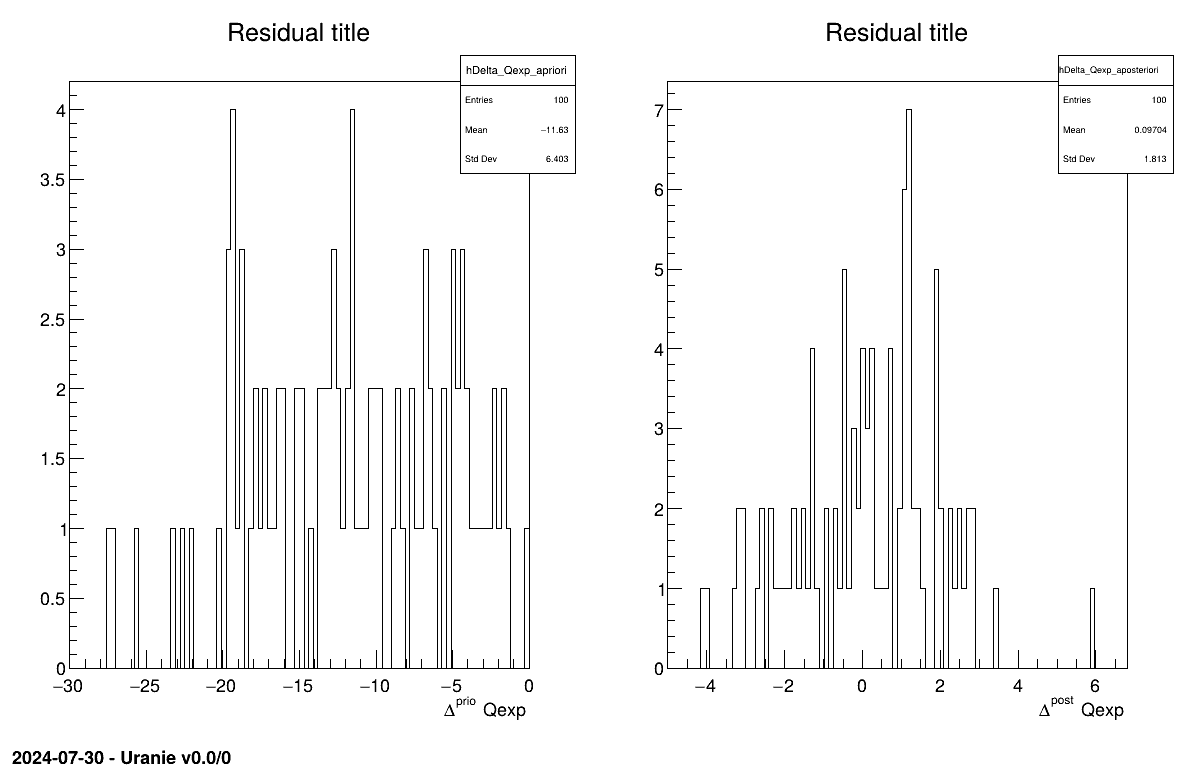
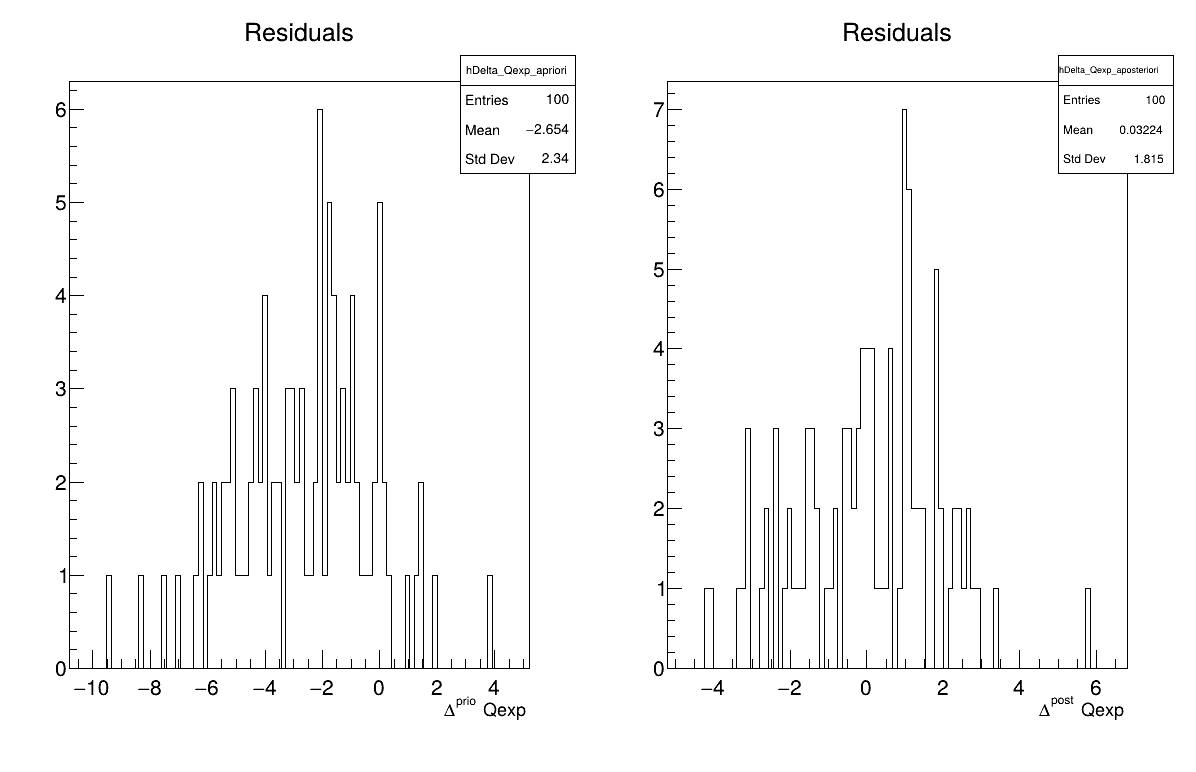
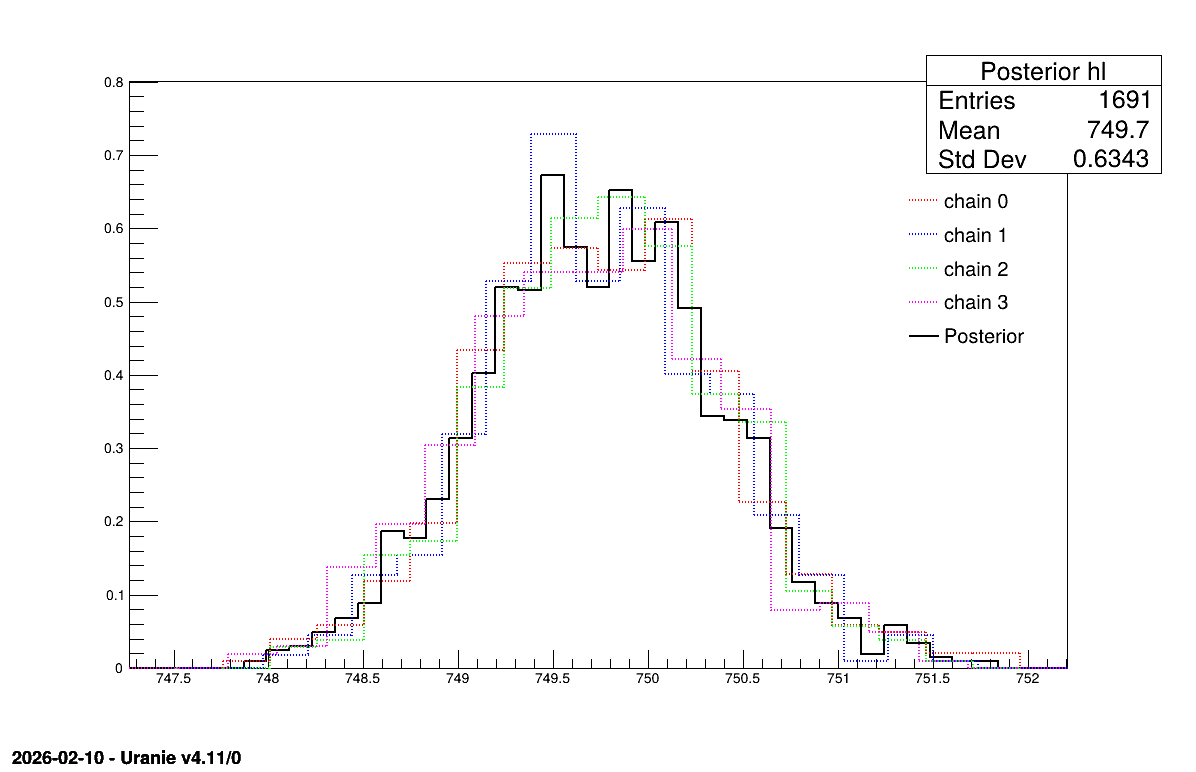
The final part demonstrates how to display the results. Since this method produces normal a posteriori distributions, they are represented by a vector of means and a covariance structure, both easily accessible. The means are displayed on screen, as illustrated in Section XIV.11.3.3. Two additional pieces of a posteriori information are presented as plots: the residuals (Figure XIV.100), which show the expected normal distribution behavior, as discussed in [metho] and the posterior distribution (Figure XIV.101).
Processing calibrationLinBayesFlowrate1D.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
*** TLinearBayesian::computeParameters(); Transformed parameters estimated are
1x1 matrix is as follows
| 0 |
------------------
0 | 749.7
The goal here is to calibrate the parameter 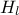 within the
within the flowrate model, while varying only two inputs
(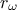 and
and
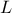 ). The remaining variables
are fixed to the following values:
). The remaining variables
are fixed to the following values: 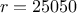 ,
, 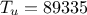 ,
, 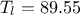 ,
, 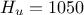 ,
, 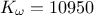 . The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the
. The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the TDataServer objects. This macro demonstrates how to apply a rejection ABC method
using a Relauncher-based architecture.
{
// Load the function flowrateCalib1D
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="Ex2DoE_n100_sd1.75.dat";
// define the reference
TDataServer *tdsRef = new TDataServer("tdsRef","doe_exp_Re_Pr");
tdsRef->fileDataRead(ExpData.Data());
// define the parameters
TDataServer *tdsPar = new TDataServer("tdsPar","pouet");
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib1D");
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
// Set the calibration object
Int_t nABC = 100; Double_t eps = 0.05;
TRejectionABC *cal = new TRejectionABC(tdsPar, runner, nABC, "");
cal->setDistance("LS",tdsRef,"rw:l","Qexp");
cal->setGaussianNoise("sd_eps");
cal->setPercentile(eps);
cal->estimateParameters();
// Compute statistics
tdsPar->computeStatistic();
cout << "The mean of hl is " << tdsPar->getAttribute("hl")->getMean() << endl;
cout << "The std of hl is " << tdsPar->getAttribute("hl")->getStd() << endl;
// Draw the parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1); padPar->Draw(); padPar->cd();
cal->drawParameters("Parameters","*","nonewcanvas");
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1); padRes->Draw(); padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
}
Much of this code has already been covered in the previous section Section XIV.11.1.2
(up to the sequential run). The main difference here is that the input parameter is now defined as a
TStochasticDistribution, representing the a priori chosen distribution.
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
The calibration object is then created by specifying both the number of elements in the final sample
(nABC = 100) and the percentile of events retained (eps = 0.05, meaning that 2000
locations will be tested). Since the code is deterministic, uncertainty is introduced by adding random Gaussian
noise. The standard deviation of this noise is defined on an event-by-event basis using a variable from the
observation dataserver. Finally, the distance function is specified, and the estimation process is performed.
// Set the calibration object
Int_t nABC = 100; Double_t eps = 0.05;
TRejectionABC *cal = new TRejectionABC(tdsPar, runner, nABC, "");
cal->setDistance("LS",tdsRef,"rw:l","Qexp");
cal->setGaussianNoise("sd_eps");
cal->setPercentile(eps);
cal->estimateParameters();
The final part demonstrates how to display the results. Since this method produces samples of the a posteriori distributions, basic statistical information are directly displayed on screen, as shown in Section XIV.11.4.3. Two additional pieces of a posteriori information are presented as plots: the residuals (Figure XIV.102), which show the expected normal distribution behavior, as discussed in [metho] and the posterior distribution (Figure XIV.103).
// Compute statistics
tdsPar->computeStatistic();
cout << "The mean of hl is " << tdsPar->getAttribute("hl")->getMean() << endl;
cout << "The std of hl is " << tdsPar->getAttribute("hl")->getStd() << endl;
// Draw the parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1); padPar->Draw(); padPar->cd();
cal->drawParameters("Parameters","*","nonewcanvas");
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1); padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
3
Processing calibrationRejectionABCFlowrate1D.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[${SOURCEDIR}/meTIER/calibration/souRCE/TDistanceLikelihoodFunction.cxx] Line[601]
<URANIE::INFO> TDistanceLikelihoodFunction::setGaussianRandomNoise: gaussian random noise(s) is added using information in [sd_eps] to modify the output variable(s) [out].
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[${SOURCEDIR}/meTIER/calibration/souRCE/TRejectionABC.cxx] Line[118]
<URANIE::INFO> TRejectionABC::computeParameters:: 2000 evaluations will be performed !
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
A posteriori mean coordinates are : (749.926)
The mean of hl is 749.926
The std of hl is 1.97042
The goal here is to calibrate the parameter 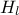 within the
within the flowrate model, while varying only two inputs
(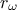 and
and
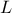 ). The remaining variables
are fixed to the following values:
). The remaining variables
are fixed to the following values: 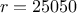 ,
, 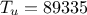 ,
, 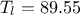 ,
, 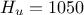 ,
, 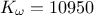 . The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the
. The context of this example has already been presented in
Section XI.2.4, including the model (implemented here as a C++ function) and the
initial lines defining the TDataServer objects. This macro demonstrates how to apply the Markov chain Monte Carlo
approach and more precisely the component-wise Metropolis-Hastings algorithm using a
Relauncher-based architecture.
{
// Load the function flowrateCalib1D
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="Ex2DoE_n100_sd1.75.dat";
// Define the reference
TDataServer *tdsRef = new TDataServer("tdsRef","doe_exp_Re_Pr");
tdsRef->fileDataRead(ExpData.Data());
// Define the parameters
TDataServer *tdsPar = new TDataServer("tdsPar","tdsPar");
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *Model = new TCIntEval("flowrateCalib1D");
Model->addInput(tdsPar->getAttribute("hl"));
Model->addInput(tdsRef->getAttribute("rw"));
Model->addInput(tdsRef->getAttribute("l"));
Model->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(Model);
// Set the calibration object
TMCMC *cal = new TMCMC(tdsPar, runner, 2000);
// Set the likelihood and reference with default gaussian log likelihood
cal->setLikelihood("log-gauss",tdsRef,"rw:l","Qexp","sd_eps");
// Set the MCMC algorithm to component-wise Metropolis-Hastings
cal->setAlgo("MH_1D");
// Set the acceptation ratio range between 20% and 50%
cal->setAcceptationRatioRange(0.2, 0.5);
// Set the number of iterations between two information displays
cal->setNbDump(500);
// Initialise 4 chains
cal->setMultistart(4);
// Set the starting points of each chain
cal->setStartingPoints(0, {740});
cal->setStartingPoints(1, {745});
cal->setStartingPoints(2, {750});
cal->setStartingPoints(3, {755});
// Set the standard deviation of the proposal for all the chains
cal->setProposalStd(-1, {5});
// Run the MCMC algorithm
cal->estimateParameters();
// Quality assessment: Draw the trace of the Markov Chain
TCanvas *canTr = new TCanvas("CanTr","CanTr",1200,800);
TPad *padTr = new TPad("padTr","padTr",0, 0.03, 1, 1);
padTr->Draw();
padTr->cd();
cal->drawTrace("Trace", "*", "nonewcanvas");
// Quality assessment: Draw the acceptation ratio
TCanvas *canAcc = new TCanvas("CanAcc","CanAcc",1200,800);
TPad *padAcc = new TPad("padAcc","padAcc",0, 0.03, 1, 1);
padAcc->Draw();
padAcc->cd();
cal->drawAcceptationRatio("Acceptation ratio", "*", "nonewcanvas");
// Set the number of iterations to remove
int burnin = 50;
cal->setBurnin(burnin);
// Verify that the burn-in has been set correctely with Gelman-Rubin statistic
std::unordered_map<string, double> GelmanRubin_values = cal->diagGelmanRubin();
// Compute Effective Sample Size
std::unordered_map<string, vector<int>> ESS_values = cal->diagESS();
// Set the lag
int lag = 1;
cal->setLag(lag);
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals", "*", "", "nonewcanvas");
// Draw the posterior distributions
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1);
padPar->Draw();
padPar->cd();
cal->drawParameters("Posterior", "*", "nonewcanvas");
}
Much of this code has already been covered in the previous section Section XIV.11.1.2
(up to the sequential run). The main difference here is that the input parameter is now defined as a
TStochasticDistribution, representing the a priori chosen distribution.
tdsPar->addAttribute( new TUniformDistribution("hl", 700.0, 760.0) );
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
The calibration object is then created by specifying both the number of iterations of the algorithm (set to 2000). The likelihood function is defined as Gaussian, with the reference dataserver, the input and output variables, and the standard deviation of the experimental data provided as arguments. Several additional options are configured: the component-wise algorithm is selected, the acceptance range is set, the interval between two console displays is specified, the number of chains to initialise is defined, the starting points of each chain are assigned (one by one), and the standard deviation of the initial proposal distribution is set (for all chains at once). For further details on these options, see Section XI.6.2. Finally, the estimation process is performed, using the chosen MCMC algorithm (component-wise Metropolis-Hastings).
// Set the calibration object
TMCMC *cal = new TMCMC(tdsPar, runner, 2000);
// Set the likelihood and reference with default gaussian log likelihood
cal->setLikelihood("log-gauss",tdsRef,"rw:l","Qexp","sd_eps");
// Set the MCMC algorithm to component-wise Metropolis-Hastings
cal->setAlgo("MH_1D");
// Set the acceptation ratio range between 20% and 50%
cal->setAcceptationRatioRange(0.2, 0.5);
// Set the number of iterations between two information displays
cal->setNbDump(500);
// Initialise 4 chains
cal->setMultistart(4);
// Set the starting points of each chain
cal->setStartingPoints(0, {740});
cal->setStartingPoints(1, {745});
cal->setStartingPoints(2, {750});
cal->setStartingPoints(3, {755});
// Set the standard deviation of the proposal for all the chains
cal->setProposalStd(-1, {5});
// Run the MCMC algorithm
cal->estimateParameters();
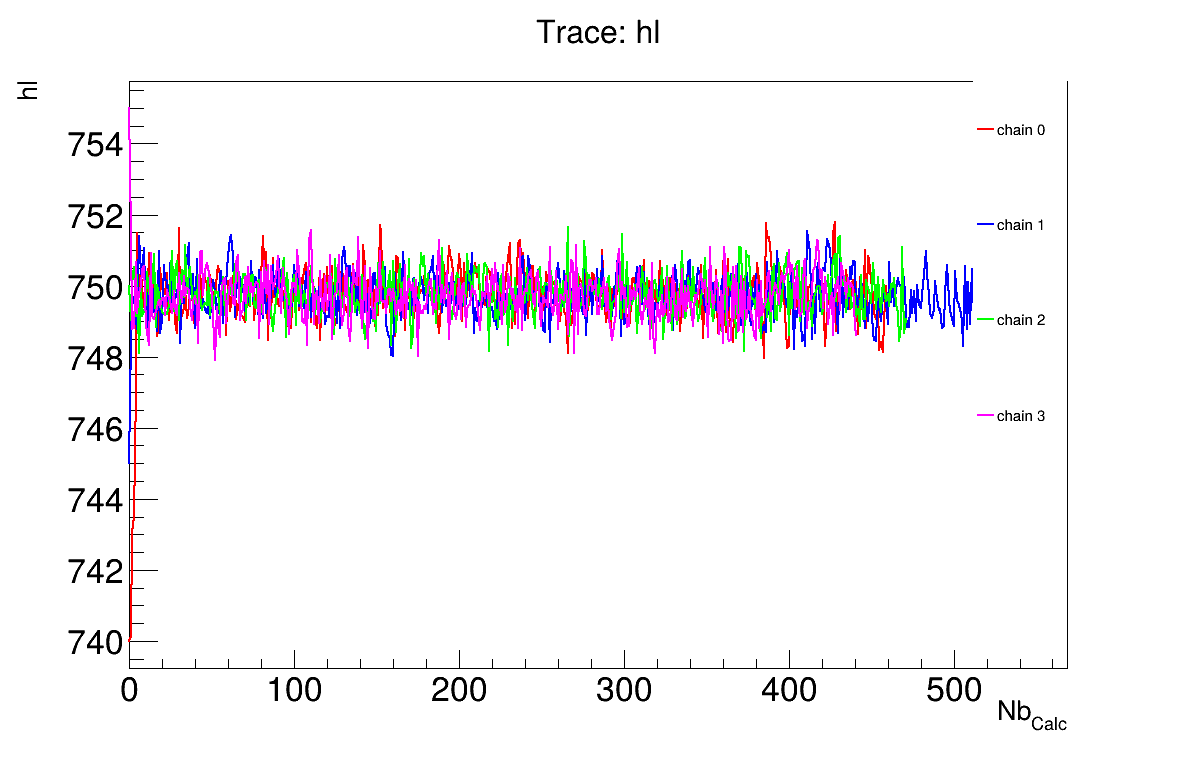
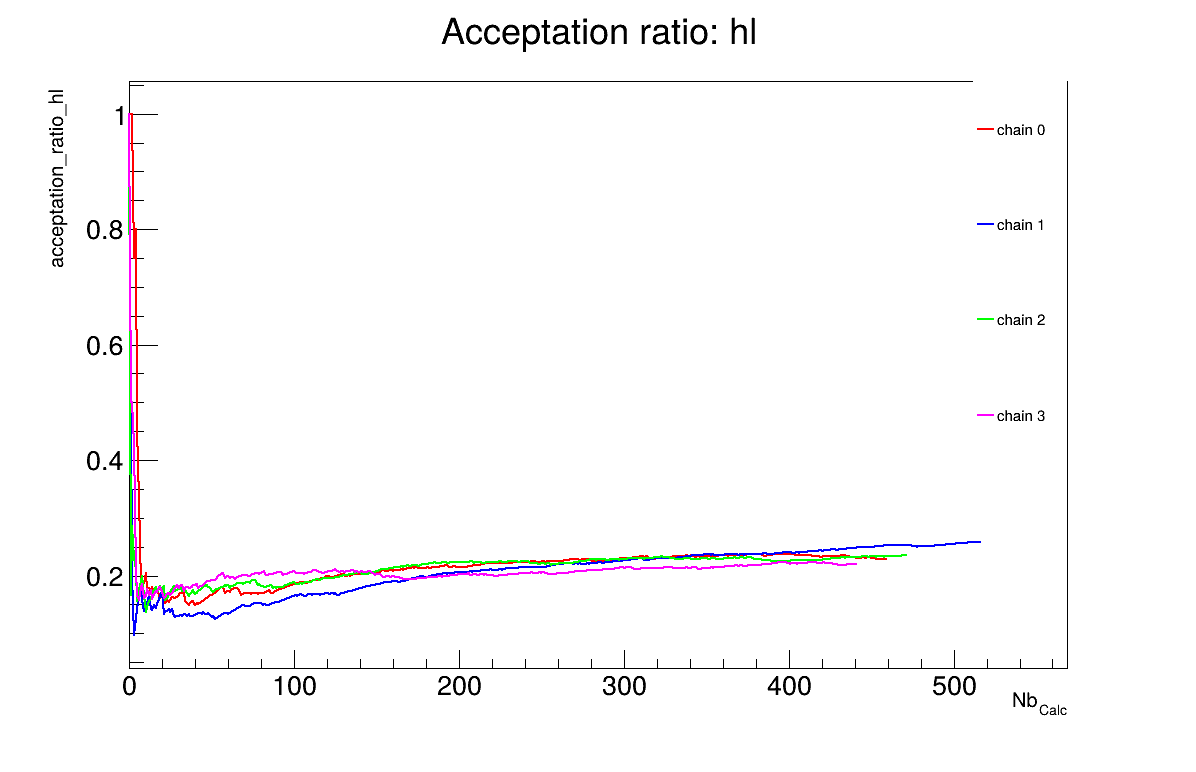
The final part demonstrates how to display and analyse the results. The first elements to examine are the trace plot (Figure XIV.104) and the acceptance ratio plot (Figure XIV.105). On these plots, you should check that the number of accepted iterations is sufficiently high (at least several hundred if the chains stabilise quickly, in our case, around 500), and that the converged acceptance ratio falls within a reasonable range (20-50%, here, it is close to 25%). When inspecting the trace, the chains should have converged around the same region (as they do here), and they should evolve rapidly, which indicates efficient exploration and low autocorrelation (also observed in our case). Finally, these plots help verify whether all chains behave stably and from which iteration they can be considered stable. This provides a first indication of a suitable burn-in value. In this example, the chains appear to have converged, and a burn-in value of 50 seems appropriate.
The burn-in size can then be defined and its consistency verified using the Gelman-Rubin statistic. In our case, a value of 1.00137 is reported in Section XIV.11.5.3 which is very close to 1 and confirms excellent convergence of the chains. Otherwise, a different burn-in value might have been more appropriate, the calculation could have been extended, or the MCMC algorithm settings adjusted.
The next step is to check for sample autocorrelation within the chains. The Effective Sample Size (ESS) provides a good indication of the lag to be used. In our case, the chains contain a reasonable number of independent samples (at least 257 each), and a lag of 1 is sufficient (see Section XIV.11.5.3). Otherwise, the calculation could have been extended, or the MCMC algorithm settings adjusted (especially the standard deviation of the proposal distribution).
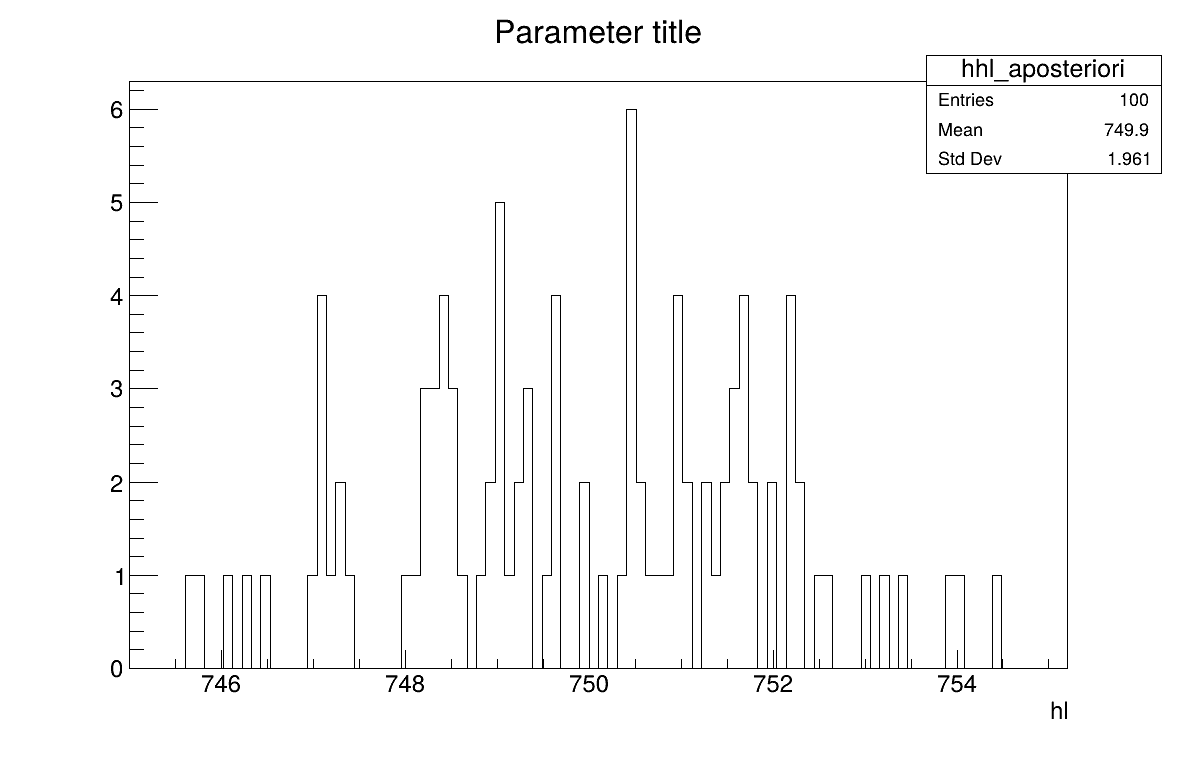
Two additional pieces of a posteriori information are presented as plots: the residuals (Figure XIV.106), which show the expected normal distribution behavior, as discussed in [metho] and the posterior distribution (Figure XIV.107).
// Quality assessment: Draw the trace of the Markov Chain
TCanvas *canTr = new TCanvas("CanTr","CanTr",1200,800);
TPad *padTr = new TPad("padTr","padTr",0, 0.03, 1, 1);
padTr->Draw();
padTr->cd();
cal->drawTrace("Trace", "*", "nonewcanvas");
// Quality assessment: Draw the acceptation ratio
TCanvas *canAcc = new TCanvas("CanAcc","CanAcc",1200,800);
TPad *padAcc = new TPad("padAcc","padAcc",0, 0.03, 1, 1);
padAcc->Draw();
padAcc->cd();
cal->drawAcceptationRatio("Acceptation ratio", "*", "nonewcanvas");
// Set the number of iterations to remove
int burnin = 50;
cal->setBurnin(burnin);
// Verify that the burn-in has been set correctely with Gelman-Rubin statistic
std::unordered_map<string, double> GelmanRubin_values = cal->diagGelmanRubin();
// Compute Effective Sample Size
std::unordered_map<string, vector<int>> ESS_values = cal->diagESS();
// Set the lag
int lag = 1;
cal->setLag(lag);
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals", "*", "", "nonewcanvas");
// Draw the posterior distributions
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1);
padPar->Draw();
padPar->cd();
cal->drawParameters("Posterior", "*", "nonewcanvas");
3
Processing calibrationMCMCFlowrate1D.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[${SOURCEDIR}/meTIER/calibration/souRCE/TMCMC.cxx] Line[193]
<URANIE::INFO> TMCMC::constructor
<URANIE::INFO> A folder named [MCMC_1] has been created to store the results of the new calculation.
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[${SOURCEDIR}/meTIER/calibration/souRCE/TMCMC.cxx] Line[673]
<URANIE::INFO> TMCMC::constructor
<URANIE::INFO> MCMC_1_chain_0 file has been duplicated in the folder [MCMC_1] to initiate [4] chains.
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [22.95%].
A posteriori mean coordinates are : (749.692)
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [25.8%].
A posteriori mean coordinates are : (749.727)
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [23.55%].
A posteriori mean coordinates are : (749.769)
500 events done
1000 events done
1500 events done
2000 events done
Computation finished with 2000 iterations.
Acceptation ratio [22.05%].
A posteriori mean coordinates are : (749.713)
Gelman-Rubin statistic for variable [hl] is equal to 1.00137. Excellent convergence.
Effective Sample Size (ESS) for variable [hl] and chain [0] is equal to 257.
Effective Sample Size (ESS) for variable [hl] and chain [1] is equal to 305.
Effective Sample Size (ESS) for variable [hl] and chain [2] is equal to 405.
Effective Sample Size (ESS) for variable [hl] and chain [3] is equal to 371.
For uncorrelated samples, the lag should be set to 1.
The goal here is to calibrate the parameters 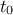 (constant term) and
(constant term) and 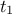 (multiplicative term) within a simple linear model (implemented in
(multiplicative term) within a simple linear model (implemented in UserFunctions.C), against the
observable outputs 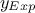 , associated
with given values of
, associated
with given values of 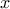 stored in
the input file
stored in
the input file linReg_Database.dat. The initial lines defining the TDataServer and the model are similar
to those presented in Section XI.2.4. This macro demonstrates how to apply the Markov chain Monte Carlo
approach and more precisely the classic Metropolis-Hastings algorithm in two dimensions using a
Relauncher-based architecture.
{
// Load the function Toy
gROOT->LoadMacro("UserFunctions.C");
// Input reference file
TString ExpData="linReg_Database.dat";
// Define the reference
TDataServer *tdsRef = new TDataServer("tdsRef","tdsRef");
tdsRef->fileDataRead(ExpData.Data());
// Define the uncertainty model guessing the standard deviation (here 0.2)
tdsRef->addAttribute("sd_eps","0.2");
// Define the parameters
TDataServer *tdsPar = new TDataServer("tdsPar","tdsPar");
double binf_search=-2.0, bsup_search=2.0;
tdsPar->addAttribute(new TUniformDistribution("t0", binf_search, bsup_search) );
tdsPar->addAttribute(new TUniformDistribution("t1", binf_search, bsup_search)) ;
// Create the output attribute
TAttribute *out = new TAttribute("out");
// Create interface to assessors
TCIntEval *eval = new TCIntEval("Toy");
eval->addInput(tdsRef->getAttribute("x"));
eval->addInput(tdsPar->getAttribute("t0"));
eval->addInput(tdsPar->getAttribute("t1"));
eval->addOutput(out);
// Set the runner
TSequentialRun *runner = new TSequentialRun(eval);
// Set the calibration object with default Metropolis-Hastings algorithm
TMCMC *cal = new TMCMC(tdsPar, runner, 4000);
// Set the likelihood and reference with default gaussian log likelihood
cal->setLikelihood("log-gauss",tdsRef,"x","yExp","sd_eps");
// Set the number of iterations between two information displays
cal->setNbDump(1000);
// Initialise 4 chains
cal->setMultistart(4);
// Set the starting points of each chain
cal->setStartingPoints(0, {0.5, -0.2});
cal->setStartingPoints(1, {-1.5, 1});
cal->setStartingPoints(2, {-1.2, -0.2});
cal->setStartingPoints(3, {0.3, 0.8});
// Set the standard deviation of the proposal for all the chains
cal->setProposalStd(-1, {0.05, 0.05});
// Run the MCMC algorithm
cal->estimateParameters();
// Quality assessment: Draw the trace of the Markov Chain
TCanvas *canTr = new TCanvas("CanTr","CanTr",1200,800);
TPad *padTr = new TPad("padTr","padTr",0, 0.03, 1, 1);
padTr->Draw();
padTr->cd();
cal->drawTrace("Trace","*","nonewcanvas");
// Quality assessment: Draw the acceptation ratio
TCanvas *canAcc = new TCanvas("CanAcc","CanAcc",1200,800);
TPad *padAcc = new TPad("padAcc","padAcc",0, 0.03, 1, 1);
padAcc->Draw();
padAcc->cd();
cal->drawAcceptationRatio("Acceptation ratio","*","nonewcanvas");
// Set the number of iterations to remove
int burnin=100;
cal->setBurnin(burnin);
// Verify that the burn-in has been set correctely with Gelman-Rubin statistic
std::unordered_map<string, double> GelmanRubin_values = cal->diagGelmanRubin();
// Compute Effective Sample Size
std::unordered_map<string, vector<int>> ESS_values = cal->diagESS();
// Set the lag
int lag=3;
cal->setLag(lag);
// Quality assessment: Draw the 2D trace of the Markov Chain
TCanvas *canTr2D = new TCanvas("CanTr2D", "CanTr2D", 1200, 800);
TPad *padTr2D = new TPad("padTr2D", "padTr2D", 0, 0.03, 1, 1);
padTr2D->Draw();
padTr2D->cd();
cal->draw2DTrace("Trace 2D", "t0:t1", "nonewcanvas");
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
// Draw the parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1);
padPar->Draw();
padPar->cd();
cal->drawParameters("Posterior","*","nonewcanvas");
}
Much of this code has already been covered in the previous section Section XIV.11.1
(up to the sequential run). The main difference here is that the input parameter is now defined as a
TStochasticDistribution, representing the a priori chosen distribution.
Another difference from the previous example Section XIV.11.5 is that no uncertainty
associated with the observables is provided in the input file. To this end, a new attribute is defined and added to the
reference dataserver, with an assumed standard deviation of 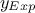 equal to 0.2.
equal to 0.2.
// Define the uncertainty model guessing the standard deviation (here 0.2)
tdsRef->addAttribute("sd_eps","0.2");
// Define the parameters
double binf_search=-2.0, bsup_search=2.0;
tdsPar->addAttribute(new TUniformDistribution("t0", binf_search, bsup_search) );
tdsPar->addAttribute(new TUniformDistribution("t1", binf_search, bsup_search)) ;
The model is defined (from a TCIntEval instance with the three input variables discussed above,
in the correct order) along with the computation distribution method (sequential).
The calibration object is then created by specifying both the number of iterations of the algorithm (set to 4000). The likelihood function is defined as Gaussian, with the reference dataserver, the input and output variables, and the assumed standard deviation of the experimental data (see previous paragraph) provided as arguments. Several additional options are configured: the interval between two console displays is specified, the number of chains to initialise is defined, the starting points of each chain are assigned (one by one), and the standard deviation of the initial proposal distribution is set (for all chains at once). For further details on these options, see Section XI.6.2. Finally, the estimation process is performed, using the default MCMC algorithm (classic Metropolis-Hastings).
// Set the calibration object with default Metropolis-Hastings algorithm
TMCMC *cal = new TMCMC(tdsPar, runner, 4000);
// Set the likelihood and reference with default gaussian log likelihood
cal->setLikelihood("log-gauss",tdsRef,"x","yExp","sd_eps");
// Set the number of iterations between two information displays
cal->setNbDump(1000);
// Initialise 4 chains
cal->setMultistart(4);
// Set the starting points of each chain
cal->setStartingPoints(0, {0.5, -0.2});
cal->setStartingPoints(1, {-1.5, 1});
cal->setStartingPoints(2, {-1.2, -0.2});
cal->setStartingPoints(3, {0.3, 0.8});
// Set the standard deviation of the proposal for all the chains
cal->setProposalStd(-1, {0.05, 0.05});
// Run the MCMC algorithm
cal->estimateParameters();
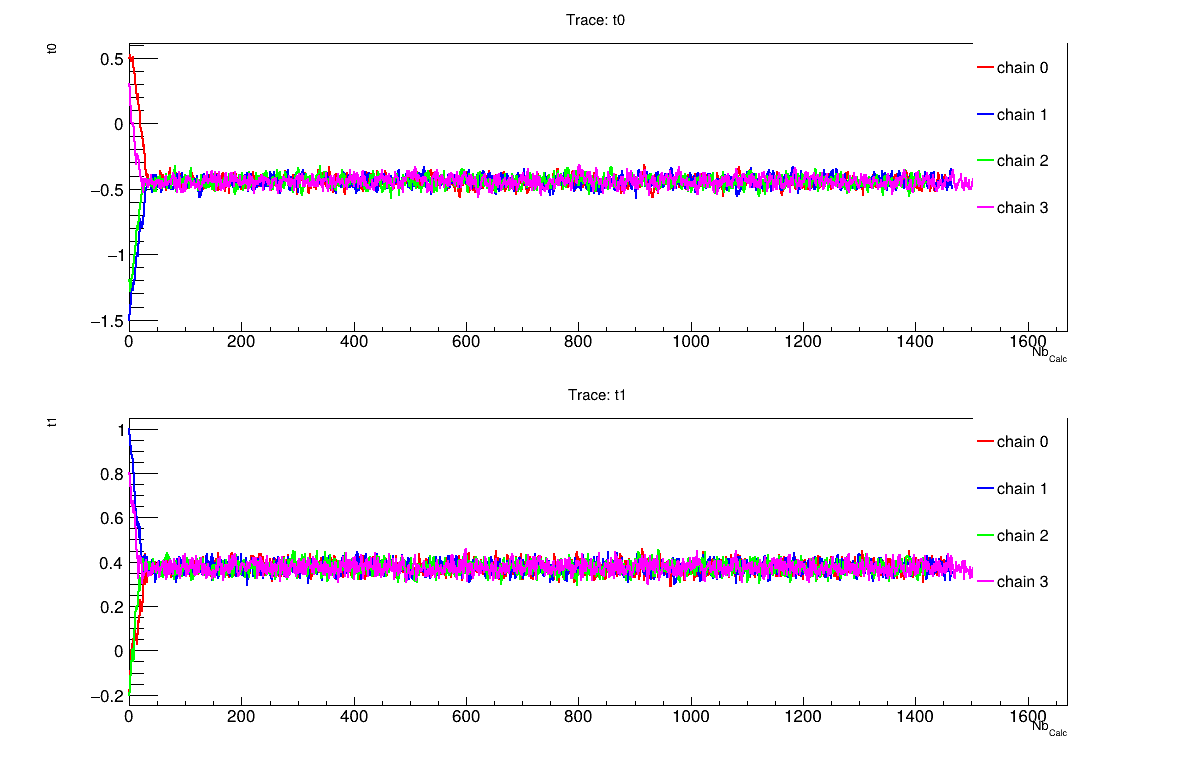
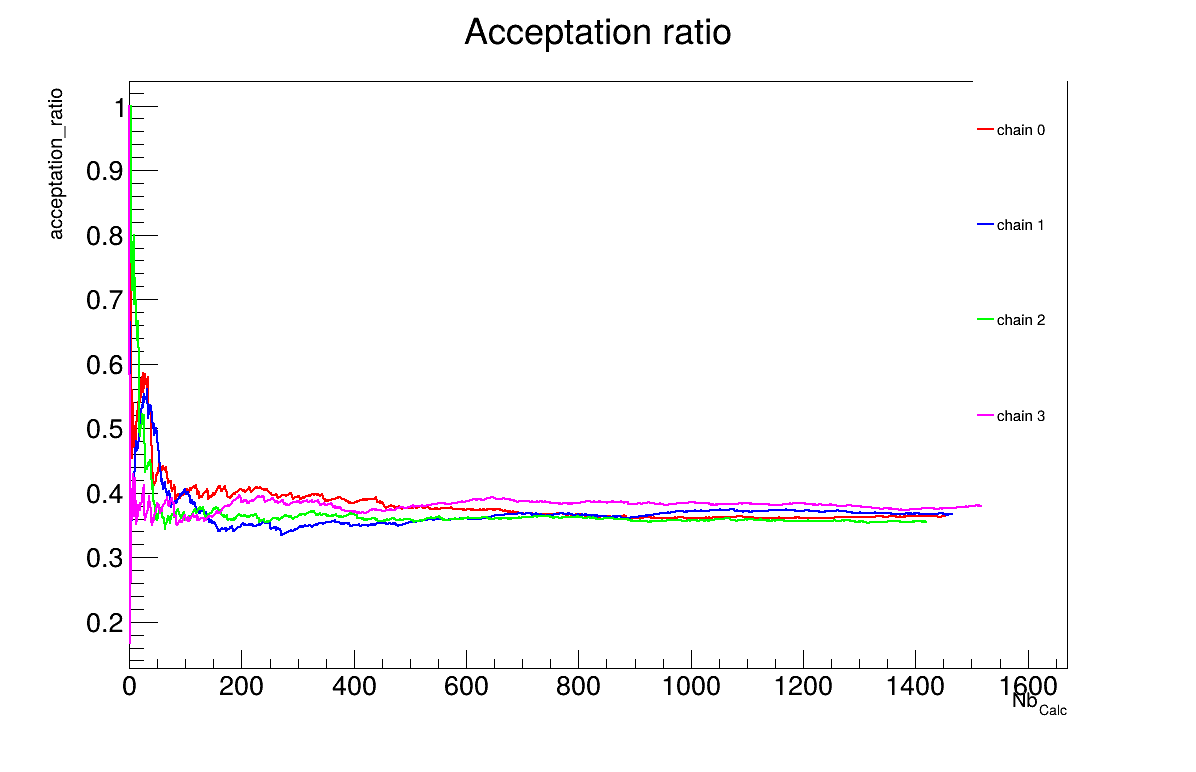
The final part demonstrates how to display and analyse the results. The first elements to examine are the trace plot (Figure XIV.108) and the acceptance ratio plot (Figure XIV.109). On these plots, you should check that the number of accepted iterations is sufficiently high (at least several hundred if the chains stabilise quickly, in our case, around 1500), and that the converged acceptance ratio falls within a reasonable range (20-50%, here, it is close to 40%). When inspecting the trace, the chains should have converged around the same region (as they do here), and they should evolve rapidly, which indicates efficient exploration and low autocorrelation (also observed in our case). Finally, these plots help verify whether all chains behave stably and from which iteration they can be considered stable. This provides a first indication of a suitable burn-in value. In this example, the chains appear to have converged, and a burn-in value of 100 seems appropriate.
The burn-in size can then be defined and its consistency verified using the Gelman-Rubin statistic. In our case,
values of 1.00092 and 0.99993 (for parameters 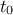 and
and 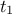 respectively)
are reported in Section XIV.11.6.3 which are very close to 1 and confirms
excellent convergence of the chains for both parameters. Otherwise, a different burn-in value might have been more
appropriate, the calculation could have been extended, or the MCMC algorithm settings adjusted.
respectively)
are reported in Section XIV.11.6.3 which are very close to 1 and confirms
excellent convergence of the chains for both parameters. Otherwise, a different burn-in value might have been more
appropriate, the calculation could have been extended, or the MCMC algorithm settings adjusted.
The next step is to check for sample autocorrelation within the chains. The Effective Sample Size (ESS) provides a good indication of the lag to be used. In our case, the chains contain a reasonable number of independent samples (at least 424 each), and a lag of 3 is sufficient (see Section XIV.11.6.3). Otherwise, the calculation could have been extended, or the MCMC algorithm settings adjusted (especially the standard deviation of the proposal distribution).
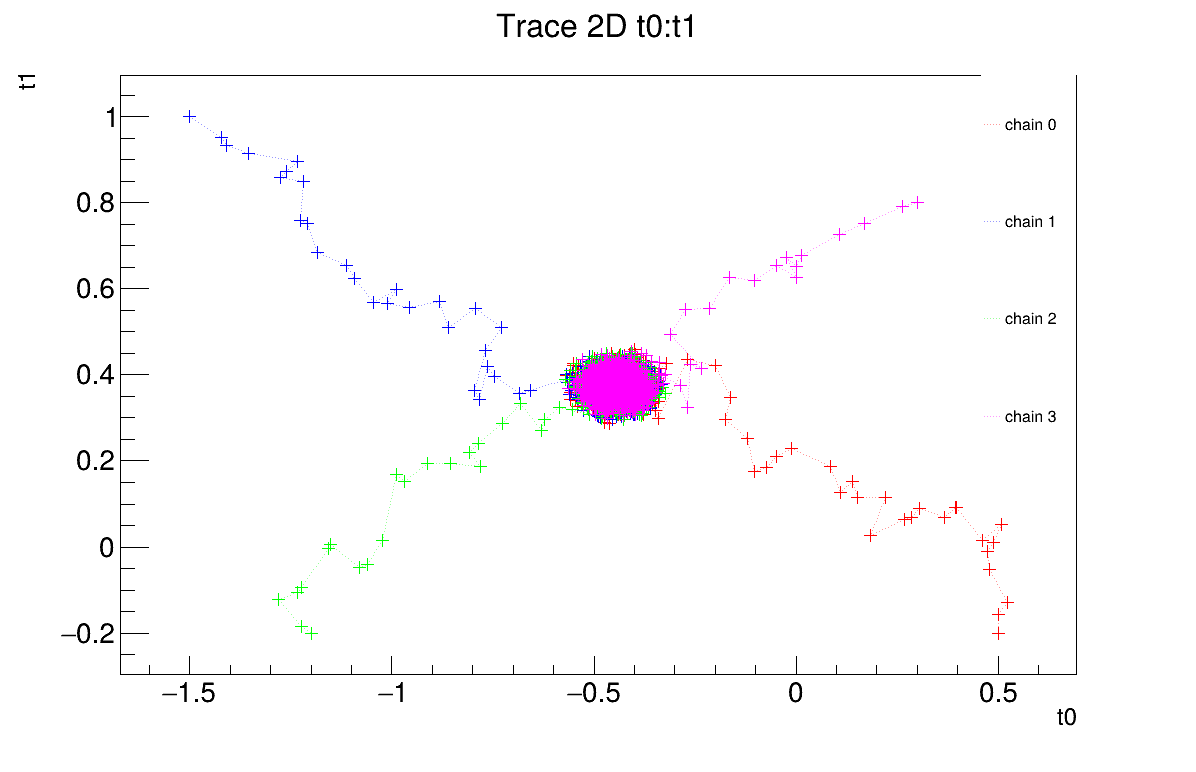
It is also possible to represent the chains in two dimensions (Figure XIV.110). This provides an alternative way to verify whether the chains converge to the same region and highlights potential covariance between the posterior distributions of the two parameters.
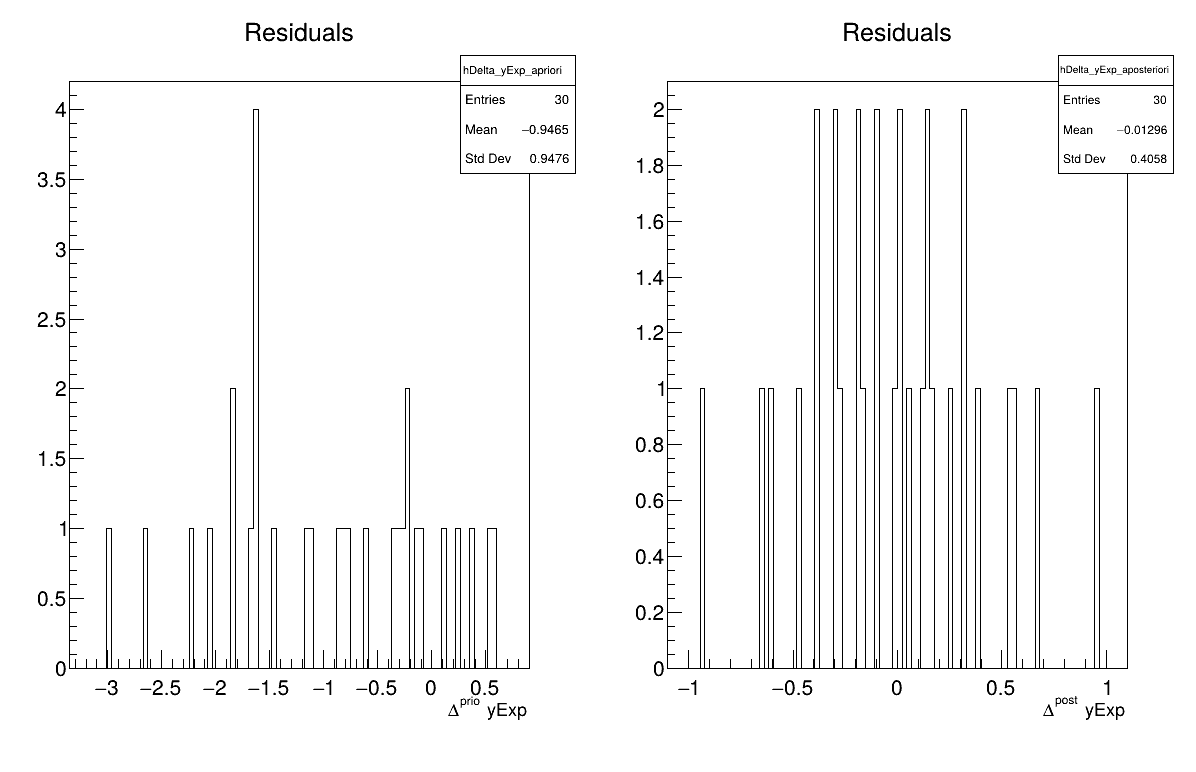
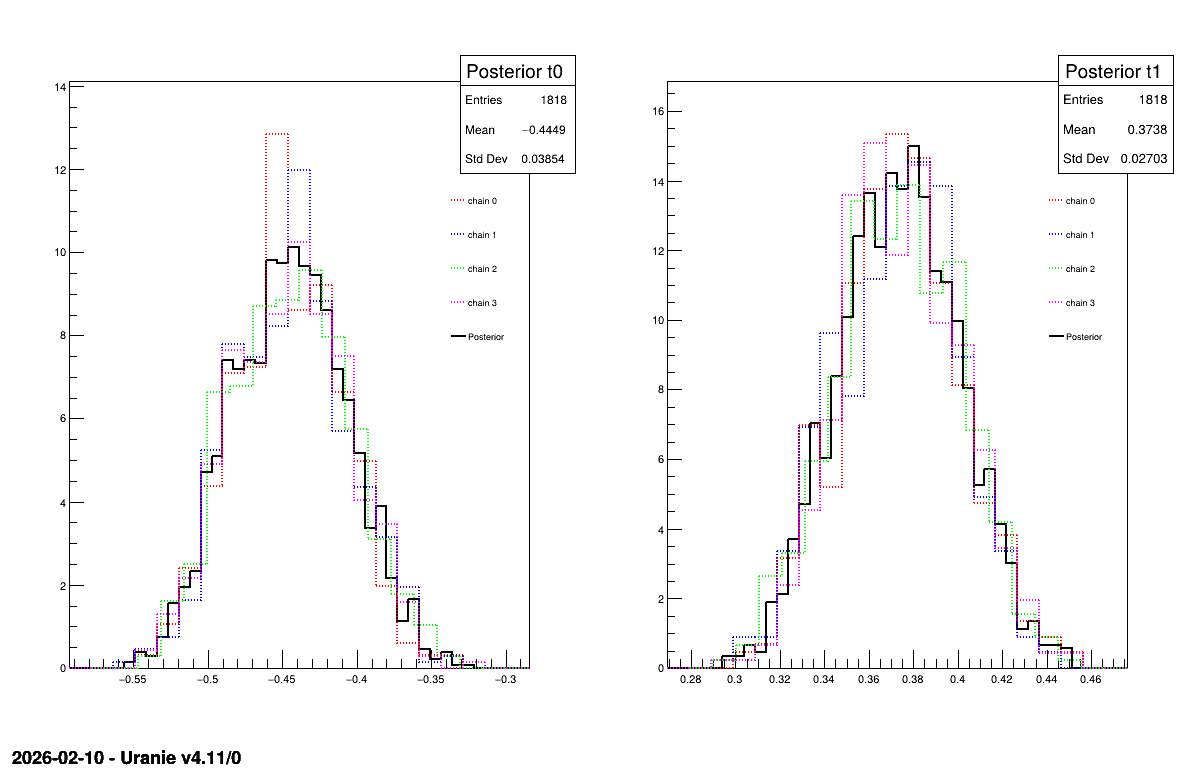
Two additional pieces of a posteriori information are presented as plots: the residuals (Figure XIV.111), which show the expected normal distribution behavior, as discussed in [metho] and the posterior distribution (Figure XIV.112).
// Quality assessment: Draw the trace of the Markov Chain
TCanvas *canTr = new TCanvas("CanTr","CanTr",1200,800);
TPad *padTr = new TPad("padTr","padTr",0, 0.03, 1, 1);
padTr->Draw();
padTr->cd();
cal->drawTrace("Trace","*","nonewcanvas");
// Quality assessment: Draw the acceptation ratio
TCanvas *canAcc = new TCanvas("CanAcc","CanAcc",1200,800);
TPad *padAcc = new TPad("padAcc","padAcc",0, 0.03, 1, 1);
padAcc->Draw();
padAcc->cd();
cal->drawAcceptationRatio("Acceptation ratio","*","nonewcanvas");
// Set the number of iterations to remove
int burnin=100;
cal->setBurnin(burnin);
// Verify that the burn-in has been set correctely with Gelman-Rubin statistic
std::unordered_map<string, double> GelmanRubin_values = cal->diagGelmanRubin();
// Compute Effective Sample Size
std::unordered_map<string, vector<int>> ESS_values = cal->diagESS();
// Set the lag
int lag=3;
cal->setLag(lag);
// Quality assessment: Draw the 2D trace of the Markov Chain
TCanvas *canTr2D = new TCanvas("CanTr2D", "CanTr2D", 1200, 800);
TPad *padTr2D = new TPad("padTr2D", "padTr2D", 0, 0.03, 1, 1);
padTr2D->Draw();
padTr2D->cd();
cal->draw2DTrace("Trace 2D", "t0:t1", "nonewcanvas");
// Draw the residuals
TCanvas *canRes = new TCanvas("CanRes","CanRes",1200,800);
TPad *padRes = new TPad("padRes","padRes",0, 0.03, 1, 1);
padRes->Draw();
padRes->cd();
cal->drawResiduals("Residuals","*","","nonewcanvas");
// Draw the parameters
TCanvas *canPar = new TCanvas("CanPar","CanPar",1200,800);
TPad *padPar = new TPad("padPar","padPar",0, 0.03, 1, 1);
padPar->Draw();
padPar->cd();
cal->drawParameters("Posterior","*","nonewcanvas");
3
Processing calibrationMCMCLinReg.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[${SOURCEDIR}/meTIER/calibration/souRCE/TMCMC.cxx] Line[193]
<URANIE::INFO> TMCMC::constructor
<URANIE::INFO> A folder named [MCMC_1] has been created to store the results of the new calculation.
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
<URANIE::INFO>
<URANIE::INFO> *** URANIE INFORMATION ***
<URANIE::INFO> *** File[${SOURCEDIR}/meTIER/calibration/souRCE/TMCMC.cxx] Line[673]
<URANIE::INFO> TMCMC::constructor
<URANIE::INFO> MCMC_1_chain_0 file has been duplicated in the folder [MCMC_1] to initiate [4] chains.
<URANIE::INFO> *** END of URANIE INFORMATION ***
<URANIE::INFO>
1000 events done
2000 events done
3000 events done
4000 events done
Computation finished with 4000 iterations.
Acceptation ratio [36.325%].
A posteriori mean coordinates are : (-0.433531,0.369056)
1000 events done
2000 events done
3000 events done
4000 events done
Computation finished with 4000 iterations.
Acceptation ratio [36.625%].
A posteriori mean coordinates are : (-0.455715,0.378325)
1000 events done
2000 events done
3000 events done
4000 events done
Computation finished with 4000 iterations.
Acceptation ratio [35.475%].
A posteriori mean coordinates are : (-0.452614,0.368962)
1000 events done
2000 events done
3000 events done
4000 events done
Computation finished with 4000 iterations.
Acceptation ratio [37.925%].
A posteriori mean coordinates are : (-0.440476,0.376135)
Gelman-Rubin statistic for variable [t0] is equal to 1.00092. Excellent convergence.
Gelman-Rubin statistic for variable [t1] is equal to 0.99993. Excellent convergence.
Effective Sample Size (ESS) for variable [t0] and chain [0] is equal to 426.
Effective Sample Size (ESS) for variable [t1] and chain [0] is equal to 893.
Effective Sample Size (ESS) for variable [t0] and chain [1] is equal to 424.
Effective Sample Size (ESS) for variable [t1] and chain [1] is equal to 727.
Effective Sample Size (ESS) for variable [t0] and chain [2] is equal to 471.
Effective Sample Size (ESS) for variable [t1] and chain [2] is equal to 594.
Effective Sample Size (ESS) for variable [t0] and chain [3] is equal to 440.
Effective Sample Size (ESS) for variable [t1] and chain [3] is equal to 802.
For uncorrelated samples, the lag should be set to 3.
The goal here is to model the uncertainties and quantify the discrepancy between model outputs and experimental data
using the Circe method on the dataset "jdd_circe_summerschool2006_dataserver.dat".
This dataset contains 150 patterns, each described by four attributes:
"exp": name of the experimental attribute;
"code": name of the code output attribute;
"sens1": name of the attribute corresponding to the derivative of the code output with respect to the first parameter;
"sens2": name of the attribute corresponding to the derivative of the code output with respect to the second parameter.
#COLUMN_NAMES: code | exp | sens1 | sens2
0.853828 0.720995 1.280695 0.426961
1.420676 1.467705 2.130798 0.710554
1.986837 1.277730 2.979664 0.994010
2.552036 1.991193 3.826800 1.277273
3.116001 2.036849 4.671714 1.560289
3.678459 3.445518 5.513915 1.843002
4.239138 4.735902 6.352916 2.125359
4.797768 3.381548 7.188232 2.407304
5.354081 5.383797 8.019378 2.688784
5.907808 5.001590 8.845874 2.969742
6.458685 5.330333 9.667245 3.250125
7.006447 7.952286 10.483015 3.529879
7.550833 4.561176 11.292717 3.808950
8.091583 7.968353 12.095884 4.087283
8.628440 8.644601 12.892057 4.364824
9.161150 7.772117 13.680780 4.641520
9.689460 11.946291 14.461602 4.917317
10.213121 9.110840 15.234080 5.192162
10.731888 9.179312 15.997775 5.466002
11.245518 12.044400 16.752254 5.738783
11.753772 9.955391 17.497091 6.010453
12.256413 8.516376 18.231867 6.280959
12.753210 11.832538 18.956171 6.550249
13.243933 15.764511 19.669597 6.818270
13.728360 13.636206 20.371749 7.084971
14.206269 15.828666 21.062237 7.350300
14.677444 15.371526 21.740682 7.614206
15.141674 17.624294 22.406711 7.876637
15.598751 16.365027 23.059960 8.137543
16.048474 13.622278 23.700075 8.396873
16.490644 16.654472 24.326711 8.654577
16.925069 18.445503 24.939533 8.910605
17.351561 21.030571 25.538214 9.164908
17.769937 17.357715 26.122438 9.417436
18.180020 11.956423 26.691900 9.668140
18.581638 19.157803 27.246304 9.916972
18.974624 16.126637 27.785365 10.163884
19.358818 18.922050 28.308810 10.408827
19.734065 20.848071 28.816374 10.651755
20.100213 18.048485 29.307807 10.892620
20.457121 8.858695 29.782866 11.131376
20.804650 17.677597 30.241324 11.367976
21.142669 22.920734 30.682962 11.602375
21.471051 18.370473 31.107575 11.834528
21.789678 16.656787 31.514968 12.064388
22.098436 19.602911 31.904959 12.291912
22.397218 27.669603 32.277379 12.517056
22.685923 22.298174 32.632070 12.739777
22.964458 21.384184 32.968887 12.960030
23.232734 30.519337 33.287695 13.177773
23.490671 15.001831 33.588376 13.392965
23.738192 25.366767 33.870822 13.605563
23.975231 23.949371 34.134936 13.815527
24.201726 13.391711 34.380637 14.022815
24.417621 21.553269 34.607854 14.227387
24.622868 26.531200 34.816530 14.429205
24.817425 20.392323 35.006621 14.628229
25.001258 32.189940 35.178096 14.824419
25.174337 20.032718 35.330935 15.017740
25.336642 26.195740 35.465133 15.208152
25.488157 30.414972 35.580695 15.395619
25.628873 29.119988 35.677642 15.580104
25.758789 32.045293 35.756006 15.761573
25.877910 21.195366 35.815830 15.939990
25.986247 28.350611 35.857174 16.115319
26.083817 38.814487 35.880105 16.287529
26.170646 31.542012 35.884708 16.456584
26.246764 26.275016 35.871075 16.622452
26.312208 38.313217 35.839315 16.785102
26.367023 28.568492 35.789546 16.944501
26.411259 31.091609 35.721899 17.100619
26.444972 16.263460 35.636518 17.253425
26.468224 21.124831 35.533558 17.402891
26.481085 33.708891 35.413184 17.548986
26.483629 24.250273 35.275576 17.691683
26.475938 35.046525 35.120922 17.830954
26.458098 24.052985 34.949423 17.966773
26.430201 34.135678 34.761291 18.099112
26.392347 27.700029 34.556749 18.227946
26.344640 32.737134 34.336029 18.353251
26.287189 28.450629 34.099376 18.475001
26.220109 23.829647 33.847044 18.593174
26.143522 27.528282 33.579297 18.707747
26.057552 36.671849 33.296408 18.818696
25.962331 26.930743 32.998661 18.926002
25.857996 32.444448 32.686349 19.029643
25.744687 28.236267 32.359775 19.129598
25.622549 22.031751 32.019249 19.225849
25.491734 21.495018 31.665091 19.318378
25.352397 21.132186 31.297629 19.407165
25.204696 19.217357 30.917198 19.492194
25.048797 26.490158 30.524145 19.573449
24.884866 21.316733 30.118819 19.650913
24.713076 25.592418 29.701580 19.724572
24.533603 19.577027 29.272793 19.794412
24.346625 34.922599 28.832833 19.860418
24.152327 35.735641 28.382076 19.922578
23.950895 19.302306 27.920910 19.980881
23.742519 22.246825 27.449724 20.035314
23.527392 27.026746 26.968916 20.085868
23.305709 22.656735 26.478887 20.132532
23.077671 15.588268 25.980044 20.175297
22.843477 26.051802 25.472798 20.214156
22.603332 26.182246 24.957565 20.249100
22.357443 38.381341 24.434763 20.280123
22.106018 22.768312 23.904816 20.307219
21.849267 26.030200 23.368151 20.330382
21.587402 15.635641 22.825196 20.349609
21.320639 18.753400 22.276382 20.364895
21.049191 20.947923 21.722145 20.376237
20.773276 24.642890 21.162919 20.383634
20.493113 14.828323 20.599142 20.387083
20.208919 17.455233 20.031254 20.386585
19.920916 16.168686 19.459693 20.382139
19.629322 15.843812 18.884899 20.373745
19.334360 18.983964 18.307313 20.361407
19.036251 20.541221 17.727376 20.345126
18.735215 14.704089 17.145526 20.324904
18.431475 27.724483 16.562203 20.300747
18.125252 20.854847 15.977844 20.272659
17.816766 15.890789 15.392886 20.240646
17.506238 16.750030 14.807764 20.204712
17.193888 10.107342 14.222909 20.164866
16.879934 15.763192 13.638752 20.121116
16.564594 27.922521 13.055719 20.073469
16.248084 14.705787 12.474234 20.021934
15.930621 9.417969 11.894719 19.966523
15.612417 13.446139 11.317589 19.907245
15.293685 13.532392 10.743257 19.844112
14.974634 20.769703 10.172131 19.777137
14.655473 17.227635 9.604615 19.706331
14.336409 12.008431 9.041108 19.631710
14.017644 17.118440 8.482001 19.553287
13.699381 14.424335 7.927684 19.471078
13.381817 14.105580 7.378537 19.385098
13.065150 8.072573 6.834935 19.295364
12.749572 10.533065 6.297249 19.201894
12.435272 11.022880 5.765839 19.104706
12.122439 12.865744 5.241061 19.003818
11.811257 10.789248 4.723263 18.899250
11.501904 5.134292 4.212786 18.791022
11.194558 11.284947 3.709961 18.679155
10.889392 10.426379 3.215113 18.563671
10.586576 8.555081 2.728559 18.444593
10.286274 10.517881 2.250605 18.321943
9.988648 12.657006 1.781552 18.195744
9.693856 9.822359 1.321689 18.066023
9.402049 9.082523 0.871296 17.932802
9.113378 12.983850 0.430646 17.796110
8.827985 6.205202 -2.89e-15 17.655971
After loading the reference dataset, the TCirce object can be constructed by specifying the reference
TDataServer the name of the experimental attribute, the code output attribute, and the derivative attributes. It is then possible
to set optional algorithm parameters before launching the process. Finally, the results are extracted and displayed in
Section XIV.11.7.3.
{
TDataServer *tds = new TDataServer();
tds->fileDataRead("jdd_circe_summerschool2006_dataserver.dat");
// tds->addAttribute("uexp", "0.05*exp");
// tds->addAttribute("sens3", "sens1*sens2");
// Create the TCirce object from the TDS and specify Experimental attribute, Code attribute and sensitivity attributes
TCirce * tc = new TCirce(tds, "exp", "code", "sens1,sens2");
// tc->setTolerance(1e-5);
// tc->setNCMatrix(5);
// TMatrixD initCMat(2,2);
// initCMat.Zero(); initCMat(0,0) = 0.042737; initCMat(1,1) = 0.525673;
// tc->setCMatrixInitial(initCMat);
// TVectorD initBVec(2);
// initBVec(0) = -1.436394; initBVec(1) = -1.501561;
// tc->setBVectorInitial(initBVec);
tc->estimate();
// Post-treatment
TVectorD vBiais = tc->getBVector();
cout << " ************** vBiais rows[" << vBiais.GetNrows() << "]"<< endl;
vBiais.Print();
TMatrixD matC = tc->getCMatrix();
cout << " ************** matC rows[" << matC.GetNrows() << "] col [" << matC.GetNcols() << "]"<< endl;
matC.Print();
}
Processing calibrationCirce.C...
--- Uranie v4.11/0 --- Developed with ROOT (6.36.06)
Copyright (C) 2013-2026 CEA/DES
Contact: support-uranie@cea.fr
Date: Tue Feb 10, 2026
*********
** addData from an another TDS [jdd_circe_summerschool2006_dataserver]
** YStar[exp] YStarSigma[]YHat[code]
** Sensitivity Attributes[sens1 sens2]
** nparameter [sens1 sens2] size[2]
** List Of TDS size[1]
Collection name='TList', class='TList', size=1
OBJ: URANIE::DataServer::TDataServer jdd_circe_summerschool2006_dataserver _title_
** List Of Informations size[3]
Collection name='TList', class='TList', size=3
OBJ: TNamed __Circe_YStar_jdd_circe_summerschool2006_dataserver_1__ exp
OBJ: TNamed __Circe_YHat_jdd_circe_summerschool2006_dataserver_1__ code
OBJ: TNamed __Circe_Sensitivity_jdd_circe_summerschool2006_dataserver_1__ sens1,sens2
** End Of addData from an another TDS [jdd_circe_summerschool2006_dataserver]
*********
**********************************
** Begin Of Initial Matrix C [1/1]
** CIRCE HAS CONVERGED
** iter[90] ** Likelihood[-2.729559333111159]
***** Selected :: iter[0] Likelihood[-2.729559333111159]
** matrix C1
2x2 matrix is as follows
| 0 | 1 |
-------------------------------
0 | 0.01612 0
1 | 0 0.03616
** vector XM1
Vector (2) is as follows
| 1 |
------------------
0 |-0.0131705
1 |0.0106155
** End Of Initial Matrix C [1/1]
**********************************
** Residual :: Mean [-0.007054035043120376] Std[1.003324966600461]
************** vBiais rows[2]
Vector (2) is as follows
| 1 |
------------------
0 |-0.0131705
1 |0.0106155
************** matC rows[2] col [2]
2x2 matrix is as follows
| 0 | 1 |
-------------------------------
0 | 0.01612 0
1 | 0 0.03616