13.6.11. Macro “sensitivityRegressionLeveLE.py”
Warning
The levele command will be installed on your machine only if a Fortran compiler is found
13.6.11.1. Objective
The objective of this macro is to perform a SRC and SRRC measurement on the temporal use-case levele.
This use-case is an example of code that takes a dozen of entries in order to compute the evolution
of dose as a function of time. The result of every computation consists in 3 vectors: the time (always
the same value disregarding all entries), the dose called “y” and a third useless information.
13.6.11.2. Macro Uranie
"""
Example of regression analysis on the levelE code
"""
import sys
from ctypes import c_double # for ROOT version greater or equal to 6.20
import numpy as np
from URANIE import Launcher, Sampler, Sensitivity
from URANIE import DataServer as DS
import ROOT
# Create DataServer and add input attributes
tds = DS.TDataServer("tds", "levelE usecase")
tds.addAttribute(DS.TUniformDistribution("t", 100, 1000))
tds.addAttribute(DS.TLogUniformDistribution("kl", 0.001, .01))
tds.addAttribute(DS.TLogUniformDistribution("kc", 1.0e-6, 1.0e-5))
tds.addAttribute(DS.TLogUniformDistribution("v1", 1.0e-3, 1.0e-1))
tds.addAttribute(DS.TUniformDistribution("l1", 100., 500.))
tds.addAttribute(DS.TUniformDistribution("r1", 1., 5.))
tds.addAttribute(DS.TUniformDistribution("rc1", 3., 30.))
tds.addAttribute(DS.TLogUniformDistribution("v2", 1.0e-2, 1.0e-1))
tds.addAttribute(DS.TUniformDistribution("l2", 50., 200.))
tds.addAttribute(DS.TUniformDistribution("r2", 1., 5.))
tds.addAttribute(DS.TUniformDistribution("rc2", 3., 30.))
tds.addAttribute(DS.TLogUniformDistribution("w", 1.0e5, 1.0e7))
# Tell the code where to find attribute value in input file
sIn = "levelE_input_with_values_rows.in"
tds.getAttribute("t").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("kl").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("kc").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("v1").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("l1").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("r1").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("rc1").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("v2").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("l2").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("r2").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("rc2").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
tds.getAttribute("w").setFileKey(sIn, "", "%e", DS.TAttributeFileKey.kNewRow)
# Create DOE
ns = 1024
samp = Sampler.TSampling(tds, "lhs", ns)
samp.generateSample()
# How to read ouput files
out = Launcher.TOutputFileRow("_output_levelE_withRow_.dat")
# Tell the output file that attribute IS a vector and is SECOND column
out.addAttribute(DS.TAttribute("y", DS.TAttribute.kVector), 2)
# Creation of Launcher.TCode
myc = Launcher.TCode(tds, " levele 2> /dev/null")
myc.addOutputFile(out)
# Run the code
tl = Launcher.TLauncher(tds, myc)
tl.run(args.runOpt) # args.runOpt = "" or "nointermed"
# Launch Regression
tsen = Sensitivity.TRegression(tds, "t:kl:kc:v1:l1:r1:rc1:v2:l2:r2:rc2:w",
"y", "SRCSRRC")
tsen.computeIndexes()
res = tsen.getResultTuple()
# Plotting mess
tps = np.array([20000, 30000, 40000, 50000, 60000, 70000, 80000, 90000, 100000,
200000, 300000, 400000, 500000, 600000, 700000, 800000, 900000,
1e+06, 2e+06, 3e+06, 4e+06, 5e+06, 6e+06, 7e+06, 8e+06, 9e+06],
dtype=float)
colors = [1, 2, 3, 4, 6, 7, 8, 15, 30, 38, 41, 46]
c2 = ROOT.TCanvas("c2", "c2", 5, 64, 1600, 500)
pad = ROOT.TPad("pad", "pad", 0, 0.03, 1, 1)
pad.Draw()
pad.Divide(3, 1)
pad.cd(1)
ROOT.gPad.SetLogx()
ROOT.gPad.SetGrid()
mg = ROOT.TMultiGraph()
res.Draw("Value", "Inp==\"__R2__\" && Order==\"Total\" && Method==\"SRRC^2\"",
"goff")
data3 = res.GetV1()
gr3 = ROOT.TGraph(26, tps, data3)
gr3.SetMarkerColor(2)
gr3.SetLineColor(2)
gr3.SetMarkerStyle(23)
mg.Add(gr3)
res.Draw("Value", "Inp==\"__R2__\" && Order==\"Total\" && Method==\"SRC^2\"",
"goff")
data4 = res.GetV1()
gr4 = ROOT.TGraph(26, tps, data4)
gr4.SetMarkerColor(4)
gr4.SetLineColor(4)
gr4.SetMarkerStyle(23)
mg.Add(gr4)
mg.Draw("APC")
mg.GetXaxis().SetTitle("Time")
mg.GetYaxis().SetTitle("#sum Sobol")
mg.GetYaxis().SetRangeUser(0.0, 1.0)
# Legend
ROOT.gStyle.SetLegendBorderSize(0)
ROOT.gStyle.SetFillStyle(0)
lg = ROOT.TLegend(0.25, 0.7, 0.45, 0.9)
lg.AddEntry(gr4, "R2 SRC", "lp")
lg.AddEntry(gr3, "R2 SRRC", "lp")
lg.Draw()
pad.cd(2)
ROOT.gPad.SetLogx()
ROOT.gPad.SetGrid()
mg2 = ROOT.TMultiGraph()
names = ["t", "kl", "kc", "v1", "l1", "r1", "rc1",
"v2", "l2", "r2", "rc2", "w"]
src = [0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
leg = ROOT.TLegend(0.25, 0.3, 0.45, 0.89, "Cumulative contributions")
leg.SetTextSize(0.035)
for igr in range(12):
sel = "Inp==\""+names[igr]+"\""
sel += "&& Order==\"Total\" && Method==\"SRC^2\" && Algo!=\"--rho^2--\""
res.Draw("Value", sel, "goff")
data = res.GetV1()
src[igr] = ROOT.TGraph()
src[igr].SetMarkerColor(colors[igr])
src[igr].SetLineColor(colors[igr])
src[igr].SetFillColor(colors[igr])
src[igr].SetPoint(0, 0.99999999*tps[0], 0)
for ip in range(26):
x = c_double(0)
y = c_double(0) # For ROOT lower than 6.20, user ROOT.Double
if igr != 0:
src[igr-1].GetPoint(ip+1, x, y)
src[igr].SetPoint(ip+1, tps[ip], y.value+data[ip])
src[igr].SetPoint(27, tps[25]*1.000000001, 0)
leg.AddEntry(src[igr], names[igr], "f")
for igr2 in range(11, -1, -1):
mg2.Add(src[igr2])
mg2.Draw("AFL")
mg2.GetXaxis().SetTitle("Time")
mg2.GetYaxis().SetTitle("SRC^{2}")
mg2.GetYaxis().SetRangeUser(0.0, 0.3)
leg.Draw()
pad.cd(3)
ROOT.gPad.SetLogx()
ROOT.gPad.SetGrid()
mg3 = ROOT.TMultiGraph()
srrc = [0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
for igr in range(12):
sel = "Inp==\""+names[igr]+"\""
sel += "&& Order==\"Total\" && Method==\"SRRC^2\" && Algo!=\"--rho^2--\""
res.Draw("Value", sel, "goff")
data = res.GetV1()
srrc[igr] = ROOT.TGraph()
srrc[igr].SetMarkerColor(colors[igr])
srrc[igr].SetLineColor(colors[igr])
srrc[igr].SetFillColor(colors[igr])
srrc[igr].SetPoint(0, 0.99999999*tps[0], 0)
for ip in range(26):
x = c_double(0)
y = c_double(0) # For ROOT lower than 6.20, user ROOT.Double
if igr != 0:
srrc[igr-1].GetPoint(ip+1, x, y)
srrc[igr].SetPoint(ip+1, tps[ip], y.value+data[ip])
srrc[igr].SetPoint(27, tps[25]*1.000000001, 0)
srrc[igr].SetTitle(names[igr])
for igr2 in range(11, -1, -1):
mg3.Add(srrc[igr2])
# mg3.Draw("a fb l3d")
mg3.Draw("AFL")
mg3.GetXaxis().SetTitle("Time")
mg3.GetYaxis().SetTitle("SRRC^{2}")
mg3.GetYaxis().SetRangeUser(0.0, 1.0)
leg.Draw()
The levele external code is located in the bin directory of the Uranie installation.
When looking at the code and comparing it to an usual Regression estimation, the organisation is completely transparent. The only noticeable (and compulsory) thing to do is to change the default type of the attribute read at the end of the job. This is done in this line:
out.addAttribute(DS.TAttribute("y", DS.TAttribute.kVector), 2)
where the output attribute is provided, changing its nature to a vector, thanks to the second argument
of the TAttribute constructor from the default (kReal) to the desired nature (kVector). Once
this is done, this information is broadcast internally to the code that knows how to deal with this
type of attribute.
The rest of the code is the graphical part, leading to the figure below (it is provided to illustrate how to represent results).
13.6.11.3. Graph
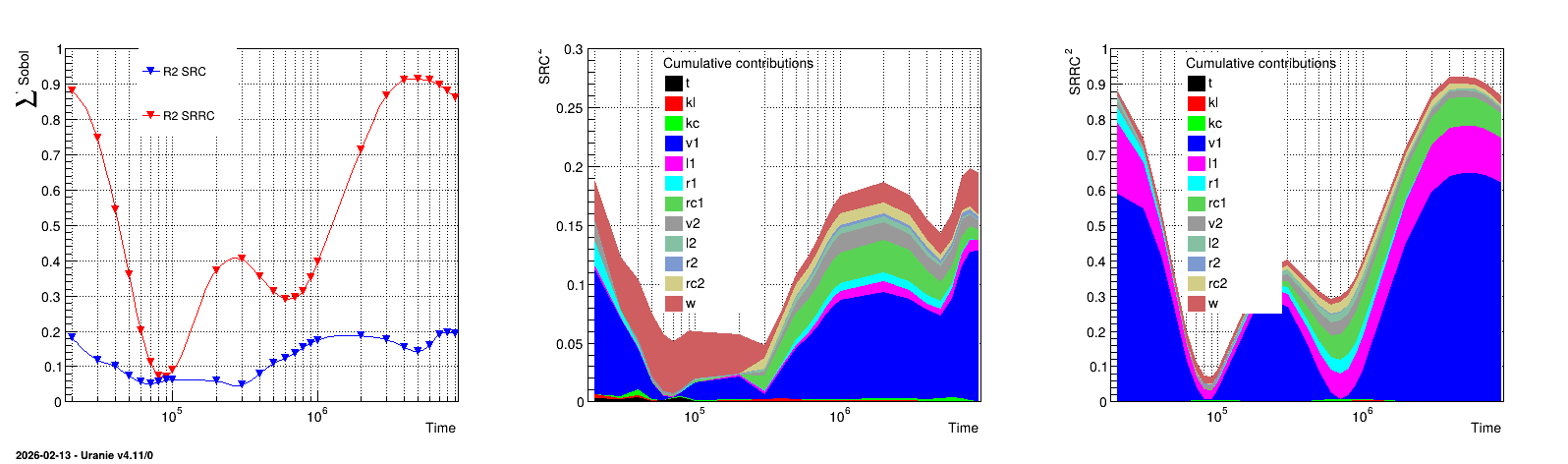
The results of the previous macro is shown in Figure 13.27, where the left panel represents the value of the \(R^2\) coefficients both the SRC and SRRC coefficients estimation. The middle and right panel display the cumulative sum of the quadratic value of the coefficient respectively for the SRC and SRRC case.
Figure 13.27 Graph of the macro “sensitivityRegressionLeveLE.py”